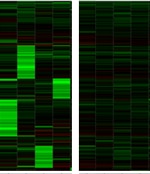
ON-TARGETplus Non-targeting Control Pool
Negative control pool of four siRNAs designed and microarray tested for minimal targeting of human, mouse or rat genes. ON-TARGETplus modifications reduce potential off-targets. Recommended for determination of baseline cellular responses in RNAi experiments.