- Gene modulation
- RNA interference
- siRNA solutions by Dharmacon™
- ON-TARGETplus 2.0 siRNA
ON-TARGETplus 2.0 siRNA
Always in Step with the Evolving Transcriptome
With Dharmacon™ ON-TARGETplus™ 2.0 siRNA, your gene-silencing reagents stay relevant to the most current transcriptome data. Our incremental realignment process ensures siRNA designs reflect the latest annotations and delivers consistent gene knockdown that is on-target while minimizing unintended off-target silencing.
How ON-TARGETplus 2.0 siRNA helps your research:
- Biologically relevant: Designs are routinely reannotated and/or redesigned according to updated genomic databases ensuring they reflect our best understanding of the genome.
- Improved specificity: By updating designs routinely ON-TARGETplus 2.0 siRNA ensures newly identified transcripts or changes to existing annotations are incorporated to minimize off-target behavior.
- Transcript isoform aware: Transcriptomics continues to rapidly advance, our refined SMARTselection™ design algorithm accounts for these changes to target as many transcripts as possible for a given gene.
Stay ahead of the evolving understanding of biology with siRNA tools that evolve with it.
1Start Here
2Choose
Knockdown target gene function with confidence
ON-TARGETplus 2.0 pre-designed siRNA is guaranteed* to knock down your gene of choice with minimal off-target activity. The next-generation RNAi SMARTselection algorithm and sequence curation used in ON-TARGETplus 2.0 siRNA strives for optimal transcriptomic coverage and further reduced off-targets by considering current gene annotations. Combined with over two decades of experience in siRNA bioinformatic design and chemical synthesis ON-TARGETplus 2.0 siRNA provides highly potent and target specific gene knockdown.
- The proprietary ON-TARGETplus™ dual strand chemical modification pattern that further minimize off-targets by up to 90% compared to unmodified siRNA (see data tab)
- Guaranteed gene silencing with our SMARTpool™ format and 3 of 4 individual siRNAs (see Guarantee tab)
- Genome-wide designs available for human, mouse, and rat genes
- Sequence information provided with every siRNA purchase
* please see guarantee tab for full guarantee details
Dharmacon ON-TARGETplus technology
Designs accounting for RNAi off-target mechanisms & contemporary transcriptomics
With our revised proprietary SMARTselection algorithm in ON-TARGETplus 2.0 siRNA our siRNA designs are built on landmark studies of seed-region–mediated off-targets and provide potent ON-TARGET knockdown and consistent experimental outcomes according to contemporary transcriptomic data to continually improve specificity and transcript isoform coverage. This is then combined with our proprietary dual-strand synthetic modification strategy to further enhance functional knockdown and minimize off-targets.
- NEW in 2.0 – Incremental transcriptome alignment & re-design: siRNAs are reannotated and re-designed according to the latest transcriptome data for improved specificity and performance. siRNAs are then curated so that individual siRNAs and SMARTpool siRNAs maximize transcript isoform coverage.
A two-stranded biological mechanism requires a two-stranded solution
Dharmacon scientists and collaborators demonstrated in 2006 (see Reference 1) that siRNA off-targets are mediated by antisense seed-region interactions, prompting the development of a dual-strand modification pattern:
- Sense strand is modified to prevent interaction with RISC and favor antisense strand uptake
- Antisense strand seed region is modified to destabilize off-target activity and enhance target specificity
Seed-region analysis on siRNA designs reduces miRNA-induced off-targets
A landmark publication (see Reference 2) from the Dharmacon research group was the first to experimentally demonstrate the key role of the seed region in mediating off-targets. A subsequent 2008 paper (see Reference 3) showed the importance of seed frequency in the 3' UTR as an indicator of its likelihood of causing off-targets. These principles were subsequently applied to our siRNA designs to improve specificity:
- Design filters exclude common seed regions likely to cause miRNA-like off-targets
- Seed frequency analysis for siRNA designs minimize off-target effects
ON-TARGETplus 2.0 benefits
- ON-TARGET knockdown – Target-specific designs with minimized off-target effects through our advanced SMARTselection design algorithm and proprietary synthetic modification patterns
- Transcriptome informed – Incrementally updated to reflect the latest human transcriptome for up to date transcript targeting
- SMARTpool siRNAs – Pools of 4 unique siRNAs further maximize on-target gene knockdown activity and minimize off-target effects while prioritizing transcript isoform coverage
- Flexible collections & libraries – Genome-wide and pathway-focused collections, as well as fully customized libraries enable screening applications with a high degree of flexibility
- Trusted experience – Dharmacon scientists are pioneers in RNAi and other gene perturbation technologies with over 25 years pioneering “rational” algorithmic design and chemical design
- Expert support – Highly trained scientific support and customer service with customer-first mindset
Product formats
- SMARTpool siRNAs: A mixture of four unique siRNAs provided as a single reagent; providing potent knockdown and further reduced off-target behavior while ensuring transcript isoform coverage.
- Set of 4: A convenient option for purchasing aliquots of all four unique individual siRNAs offered in the SMARTpool targeting a single gene for individual siRNA deconvolution.
- Individual siRNAs: For experiments where a single design has been identified for specific transcript activity or preferred performance in a given experiment.
Note: SMARTpool siRNAs and Set of 4 siRNAs are designed to contain four unique seeds and non-overlapping genomic coordinates while striving for optimal transcript coverage. See Quick Facts tab for more information.
Complete your RNAi experiment using siRNA
- ON-TARGETplus 2.0 siRNA for your gene target of choice
- Appropriate Positive and non-targeting controls
- DharmaFECT™ transfection reagent or electroporation for delivery
- siRNA buffer and/or RNase-free water for resuspension of siRNA
Order quantity guidelines
ON-TARGETplus 2.0 reagents are routinely used at 5 to 25 nM concentration. The calculations below, based on 25 nM, are for estimation purposes only and assume no loss from pipetting. ON-TARGETplus 2.0 siRNAs are guaranteed to knock down with three out of four siRNA designs and SMARTpool when used at 100 nM.
| Approximate # reactions (wells) at 25 nM siRNA concentration | |||
|---|---|---|---|
| nmol | 96-well plate (100 µL total reaction volume) | 24-well plate (500 µL total reaction volume) | 12-well plate (1000 µL total reaction volume) |
| 1 | 400 | 80 | 40 |
| 2 | 800 | 160 | 80 |
| 5 | 2000 | 400 | 200 |
| 10 | 4000 | 800 | 400 |
| 20 | 8000 | 1600 | 800 |
Custom siRNA design
Don’t see an siRNA for your given gene of interest, or for your favorite transcript? Design and order custom siRNA sequences using our siDESIGN™ Center.
For internal research use only. Not for use in diagnostic, therapeutic procedures, or commercial activities.
Our siRNA knockdown guarantee
ON-TARGETplus 2.0 siRNA reagents (SMARTpool siRNA and three out of four individual siRNAs) are guaranteed to silence target gene expression by at least 75% at the mRNA level compared to non-targeting controls as measured by qPCR when demonstrated to have been used under optimal delivery conditions (confirmed using functionally validated positive control and measured at 24 to 72 hours after transfection using 100 nM siRNA).
Note: While we guarantee knockdown >75% at 100 nM, most ON-TARGETplus 2.0 siRNA products are highly functional at 5–25 nM working concentration. Titration is recommended to maximize on-target siRNA effect and minimize any off-target behavior. Optimal siRNA delivery conditions and working concentration are experiment specific.
Please reach out to our technical support team for any questions or for guidance on optimizing the use of your siRNA products.
Complete your knockout experiment by adding these supporting reagents to your order
ON-TARGETplus 2.0 siRNA controls
Validated positive controls and non-targeting negative controls designed for use with next-generation ON-TARGETplus 2.0 siRNAs. Featuring proprietary ON-TARGETplus synthetic modifications for superior specificity and minimized off-target effects. Available as pooled or single siRNAs.
ON-TARGETplus 2.0 control siRNAs deliver improved specificity, reduced seed-based off-targeting, and enhanced consistency across experiments. These controls are recommended for any siRNA workflow utilizing ON-TARGETplus 2.0 reagents. Pooled siRNA controls are recommended when additional off-target reduction is desired and are ideal for use alongside ON-TARGETplus 2.0 pooled siRNA reagents.
Select a relevant positive control to efficiently silence a robustly expressed housekeeping gene in your experimental system for confidence in your reagent’s function.
| ON-TARGETplus 2.0 Positive Control Reagents | Species | Catalog Number |
|---|---|---|
| ON-TARGETplus 2.0 Cyclophilin B Control siRNA | Human | D2-001820-01 |
| Mouse | D2-001820-02 | |
| Rat | D2-001820-03 | |
| ON-TARGETplus 2.0 Cyclophilin B Control Pool | Human | D2-001820-10 |
| Mouse | D2-001820-20 | |
| Rat | D2-001820-30 | |
| ON-TARGETplus 2.0 GAPD Control siRNA | Human | D2-001830-01 |
| Mouse | D2-001830-02 | |
| ON-TARGETplus 2.0 GAPD Control Pool | Human | D2-001830-10 |
| Mouse | D2-001830-20 | |
| ON-TARGETplus 2.0 Negative Control Reagents | Species | Catalog Number |
| ON-TARGETplus 2.0 Non-targeting siRNAs | Human, Mouse, Rat | D2-001810-0X |
| ON-TARGETplus 2.0 Non-targeting Pool | Human, Mouse, Rat | D2-001810-10 |
Transfection reagents
DharmaFECT transfection reagents are optimized for improved delivery and reduced toxicity. The optimal DharmaFECT reagent for your experiment will depend on your cell type. consult our DharmaFECT cell type guide for recommendations on selecting the right transfection reagent.
Order quantity guidelines
ON-TARGETplus 2.0 reagents are routinely used at 5 to 25 nM concentration. The calculations below, based on 25 nM, are for estimation purposes only and assume no loss from pipetting. ON-TARGETplus 2.0 siRNAs are guaranteed to knock down with three out of four siRNA designs and SMARTpool when used at 100 nM.
| Approximate # reactions (wells) at 25 nM siRNA concentration | |||
|---|---|---|---|
| nmol | 96-well plate (100 µL total reaction volume) | 24-well plate (500 µL total reaction volume) | 12-well plate (1000 µL total reaction volume) |
| 1 | 400 | 80 | 40 |
| 2 | 800 | 160 | 80 |
| 5 | 2000 | 400 | 200 |
| 10 | 4000 | 800 | 400 |
| 20 | 8000 | 1600 | 800 |
ON-TARGETplus 2.0 and siRNA SMARTpool supporting data :
ON-TARGETplus and ON-TARGETplus 2.0 siRNAs potently deplete TP53 mRNA expression with minimal impact on global gene expression

U2OS cells were transfected with 25 nM ON-TARGETplus or ON-TARGETplus 2.0 siRNAs targeting the TP53 gene, or a non-targeting control (NTC) siRNA with 0.2 µL/well DharmaFECT 4 transfection reagent. A. After 48 hours, total RNA was isolated and prepped for RNAseq or RT-qPCR. A. Data plotted as a histogram showing the distribution of global transcriptional expression, represented as transcripts per million (TPM). Strong downregulation of TP53 mRNA expression was observed in all four siRNA treatment groups, denoted on the x-axis and represented as log2 transformed TPM fold change while global gene expression was largely unchanged relative to the corresponding NTC siRNA. n = 2 biologically independent samples per group. B. Relative gene expression was measured using RT-qPCR. The relative expression of TP53 was calculated with the ∆∆Cq method using ACTB as the housekeeping gene and normalized to an NTC. Robust target gene knockdown was observed with both ON-TARGETplus and ON-TARGETplus 2.0 siRNAs. n = 2 biologically independent samples per group.
ON-TARGETplus 2.0 siRNA achieves high knockdown efficiency with superior mitigation of off-targeting over competitor siRNA designs in U2OS cells

U2OS cells were transfected with 25 nM of four individual ON-TARGETplus 2.0 siRNAs and 25 nM of the ON-TARGETplus 2.0 siRNA SMARTpool along with three competitor siRNA designs targeting TP53 using vendor recommended concentrations and procedures. TP53 expression was measured via qPCR and number of unique off-targets associated with each condition assessed via RNAseq. All siRNAs achieved high knockdown efficiency, but two out of three competitor designs were associated with hundreds of unique off-targets. Additionally, the siRNA SMARTpool exhibited a superior off-target profile than three out of four individual ON-TARGETplus 2.0 siRNA designs as expected due to lower concentrations of each individual siRNA in the pool vs individual conditions.
Competitive benchmarking experiments highlight consistent potent target knockdown and minimal off-targeting using ON-TARGETplus 2.0 siRNA SMARTpools compared to competitor siRNA designs

U2OS cells were transfected with 25 nM of four individual ON-TARGETplus 2.0 siRNAs (not shown) and 25 nM of the ON-TARGETplus 2.0 siRNA SMARTpools along with three competitor siRNA designs targeting per target (BUB1, BRD2, and, DUSP1) using vendor recommended concentrations and procedures. Gene expression was measured via qPCR and number of unique off-targets associated with each condition assessed via RNAseq. In all experiments, ON-TARGETplus 2.0 siRNA SMARTpools produced potent gene knockdown with consistently lower unique off-targets compared to competitor designs.
ON-TARGETplus 2.0 siRNA delivers highly potent target gene knockdown similar to original ON-TARGETplus siRNA designs not impacted by transcriptomic shifts over time

U2OS or HCT-116 cells were transfected with 25 nM ON-TARGETplus or ON-TARGETplus 2.0 siRNAs targeting the AHCY (left) or PRDX1 (right) genes, or a non-targeting control (NTC) siRNA with 0.2 µL/well DharmaFECT 4 transfection reagent (U2OS) or DharmaFECT 2 transfection reagent (HCT-116). After 48 hours, total RNA was isolated and relative gene expression was measured using RT-qPCR. The relative expression of each gene was calculated with the ∆∆Cq method using ACTB as the housekeeping gene and normalized to an NTC. Robust target gene knockdown was observed with both ON-TARGETplus and ON-TARGETplus 2.0 siRNAs. n = 2 biologically independent samples per group.
ON-TARGETplus 2.0 siRNA mitigates newly discovered off-target effects arising from reference genome transcriptome expansion

U2OS or HEK293T cells were transfected with 25 nM ON-TARGETplus or ON-TARGETplus 2.0 siRNAs targeting the YY1AP1 gene, or a non-targeting control (NTC) siRNA with 0.2 µL/well DharmaFECT 1 transfection reagent (HEK293T) or DharmaFECT 4 transfection reagent (U2OS). After 48 hours, total RNA was isolated and prepped for RNAseq or RT-qPCR. A. Data plotted as a histogram showing the distribution of global transcriptional expression in U2OS cells, represented as transcripts per million (TPM). Strong downregulation of YY1AP1 mRNA expression was observed in all four siRNA treatment groups, denoted on the x-axis and represented as log2 transformed TPM fold change while global gene expression was largely unchanged relative to the corresponding NTC siRNA. Notably, ON-TARGETplus siRNAs also depleted expression of the off-target gene GON4L whereas ON-TARGETplus 2.0 siRNAs maintained high specificity and did not affect GON4L expression. n = 2 biologically independent samples per group. B. Relative gene expression was measured using RT-qPCR. The relative expression of each gene was calculated with the ∆∆Cq method using ACTB as the housekeeping gene and normalized to an NTC. Robust target gene knockdown was observed with both ON-TARGETplus and ON-TARGETplus 2.0 siRNAs, however the ON-TARGETplus siRNA was found to also reduce expression of an off-target gene, GON4L. The ON-TARGETplus 2.0 siRNAs maintained high specificity and did not affect GON4L expression. n = 2 biologically independent samples per group.
Supporting Data for ON-TARGETplus siRNA dual-strand chemical modifications, the original SMARTselection algorithm, and siRNA SMARTpool format
ON-TARGETplus siRNA chemical modifications reduce the overall number of off-targets and pooling reduces them even further

Panels (A) and (B) are representative examples of off-target signatures with and without application of ON-TARGETplus modifications to (A) a single siRNA and (B) a SMARTpool reagent. Green bars indicate genes with 2-fold or more reduction of expression when treated with the indicated siRNA reagent. The ON-TARGETplus modifications reduced the off-targets when compared to unmodified siRNA. Pooling of siRNA and the ON-TARGETplus modification pattern independently, and in combination, provide significant reduction in off-target gene silencing. Panel (C) represents quantitation of off-targets (down-regulated by 2-fold or more) induced by the indicated siRNA reagents targeting 10 different genes (4 siRNAs per gene or a single SMARTpool reagent). Off-targets were quantified using microarray analysis (Agilent), then compiled. Each shaded box represents the middle 50% of the data set. Horizontal line in box: Median value of the data set. Vertical bars: minimum and maximum data values.
False phenotypes due to off-targets are alleviated by ON-TARGETplus siRNA SMARTpool reagents while target gene knockdown is maintained

The effect of silencing ARPC1B on cell migration was studied in a breast cancer cell line. A monolayer of cells was uniformly scraped and the rate of cell migration to close the scrape (wound healing) was evaluated. Both unmodified and ON-TARGETplus siRNA reagents induced potent target knockdown. Inconsistent phenotypes due to off-target effects (red outline), were observed for cells transfected with unmodified individual siRNAs. The unmodified SMARTpool siRNAs improved the false phenotype considerably, while the ON-TARGETplus SMARTpool siRNAs significantly reduced off-target effects to produce a consistent phenotype. In collaboration with Kaylene Simpson, Laura Selfors, and Joan Brugge, Harvard Medical School.
Only the ON-TARGETplus modification pattern addresses both siRNA strands for premium silencing

The ON-TARGETplus dual-strand chemical modification begins with the sense (passenger) strand being blocked from RISC uptake to favor antisense (guide) strand loading and reduce passenger strand-induced off-targets. However, the majority of siRNA off-targets are driven by the seed region of the guide strand. ON-TARGETplus siRNA is modified within its seed region to destabilize miRNA-like activity and improve specificity to the desired target for potent knockdown.
SMARTselection algorithmic design of siRNA with low-frequency seed regions ensure fewer off-targets
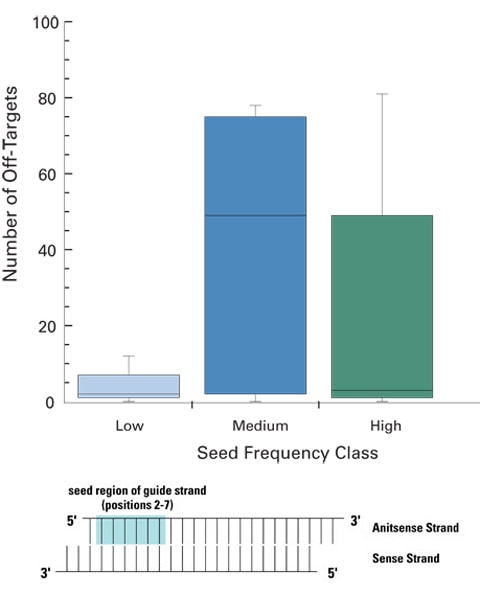
ON-TARGETplus siRNA designs leverage sophisticated bioinformatics to reduce the likelihood of miRNA-like off-targets from high-frequency or highly conserved miRNA seed regions. siRNAs with low seed frequency have a significantly lower number of off-targets than siRNAs with medium or high frequency seeds. Five siRNAs with low, medium, or high frequency seed regions were transfected into HeLa cells and their associated off-target signatures assessed via global expression profiling (Agilent 22K platform). siRNA sequences were constant at positions 1 and 8-19, only the seed regions (positions 2-7) were altered.
Low frequency seeds: < 350 occurrences in the HeLa transcriptome
Medium frequency: 2500-2800 occurrences
High frequency: >3800 occurrences
- A. L. Jackson et al., Position-specific chemical modification increases specificity of siRNA-mediated gene silencing. RNA. 12(7), 1197-1205 (July 2006).
- A. Birmingham et al., 3'-UTR seed matches, but not overall identity, are associated with RNAi off-targets. Nature Methods. 3(3), 199-204 (March 2006).
- E. M. Anderson et al., Experimental validation of the importance of seed complement frequency to siRNA specificity. RNA. 14(5), 853-861 (May 2008).
Please see our full FAQ page for all commonly asked questions about our products.
- What is the difference between ON-TARGETplus legacy siRNA and ON-TARGETplus 2.0 siRNA?
- How does ON-TARGETplus 2.0 siRNA reduce off-target effects?
- Are ON-TARGETplus 2.0 siRNAs guaranteed to knock down my target gene?
- Can I use ON-TARGETplus 2.0 siRNAs alongside my original ON-TARGETplus siRNAs and controls in an experiment?
- Can ON-TARGETplus 2.0 siRNA be used in high-throughput or automated platforms?
- Are ON-TARGETplus 2.0 siRNAs available for multiple species?
- How does ON-TARGETplus 2.0 siRNA support genome-wide and gene-family studies?
- Can ON-TARGETplus 2.0 siRNA be used alongside Edit-R™ CRISPR reagents?
- How do the four siRNAs provided in a SMARTpool siRNA format or set of 4 differ?
ON-TARGETplus 2.0 siRNA is a next-generation re-design of the original ON-TARGETplus siRNA. It features improved design algorithms, incremental transcriptome realignment, and revised sequence ranking and selection to enhance specificity through minimized off-target effects and ensure consistent gene knockdown across as many transcript isoforms as possible. Legacy ON-TARGETplus designs were based on earlier transcriptome data and design selection choices reflected earlier understanding of the transcriptomic landscape.
Each siRNA is algorithmically designed using updated transcriptomic data and designs are curated incorporating specificity as a key variable. Seed-region selection based on low frequency endogenous seeds and the ON-TARGETplus dual-strand chemical modifications further minimize unintended interactions.
Yes. ON-TARGETplus 2.0 siRNAs are backed by a gene knockdown guarantee, ensuring consistent and dependable performance when used under recommended conditions. If our siRNA fails to deliver on this guarantee, our teams will provide free replacement of alternative designs to suit your needs.
Yes, the chemical modification pattern is the same for both products, so they are suitable to be used side-by-side in an experiment. Many original ON-TARGETplus siRNA designs are highly functional and deliver excellent ON-TARGET knockdown results.
Absolutely. ON-TARGETplus 2.0 siRNAs are available in formats compatible with automated systems, including pre-arrayed or customized plates suitable for high-throughput applications.
Yes. Pre-designed ON-TARGETplus 2.0 siRNAs are available for human, mouse, and rat gene targets. For other species we recommend using our siDESIGN Center tool.
ON-TARGETplus 2.0 collections cover whole-genome libraries as well as focused sets for commonly studied gene families, enabling both broad and targeted screening approaches. In addition, custom libraries can be built according to your desired targets using the Cherry-pick tool.
Yes. ON-TARGETplus 2.0 siRNA serves as an ideal orthogonal method to validate CRISPR screening hits or complement gene knockout studies with gene knockdown testing.
The four siRNAs provided per gene are selected from a pool of algorithmically designed and curated candidates based on a number of proprietary key variables prioritizing function, specificity, and the transcript annotations for a given gene. Each set or pool of four siRNAs is selected to contain unique seeds and, wherever possible, to target the maximum number of 100% non-overlapping genomic coordinates for a given gene within design-space limitations.
Learn how we design our off-the-shelf ON-TARGETplus siRNAs and why periodic review of them is wise.
Application notes
Product inserts
Protocols
Safety data sheets
Selection guides
Related Products
Negative control pool of four siRNAs designed and microarray tested for minimal targeting of human, mouse or rat genes. ON-TARGETplus modifications reduce potential off-targets. Recommended for determination of baseline cellular responses in RNAi experiments.
Validated positive control pool of four siRNAs targeting GAPD gene in human, mouse, or rat. Includes patented ON-TARGETplus modifications to minimize off-target effects. Useful for determination of optimal RNAi conditions.
Concentrated buffer solution recommended for resuspension and long-term storage of any short, double-strand, or single-strand synthetic RNA molecule. Dilute with RNase-free water prior to use.
Catalog ID:B-002000-UB-100
Unit Size:100 mL
$99.00
An aliquot of each of the four DharmaFECT formulations for siRNA/microRNA transfection optimization studies
Catalog ID:T-2005-01
Unit Size:0.2 mL
