Add any of our CRISPR portfolio reagents (CRISPRko, CRISPRa, CRISPRi) to your cart to see new pricing (up to 56% off) now including All-in-one lentiviral sgRNA formats in 2026
CRISPR interference (CRISPRi) is a novel method for specific gene knockdown.
The CRISPRi system uses a deactivated Cas9 nuclease (dCas9) fused to our proprietary SALL1-SDS3 repressor construct to block transcription of target genes, without cutting the DNA.
CRISPRi benefits |
Ideal applications |
|
|
|
|
|
|
|
The CRISPRi system
The Dharmacon™ CRISPRmod CRISPRi system requires two components: a specialized dCas9 fused to repressor proteins (SALL1 and SDS3), and a guide RNA specifically designed to target the region immediately downstream of a gene’s transcriptional start site (TSS). The guide associates with dCas9-SALL1-SDS3 and directs the repressor complex to the DNA target site (figure 1).
We offer lentiviral and transient approaches for both repressor protein and guide introduction.
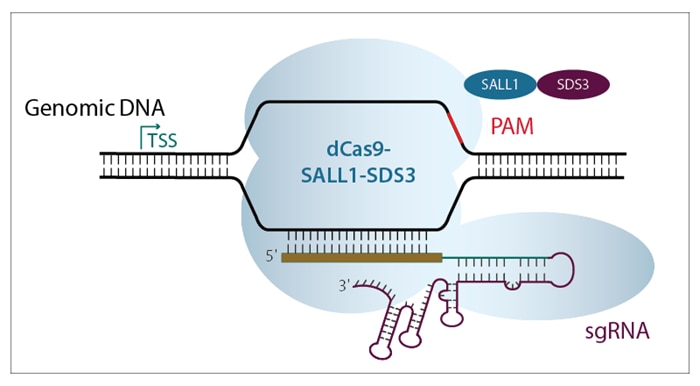
Figure 1. Diagram of dCas9-SALL1-SDS3 with sgRNA targeting the region immediately downstream of a gene’s transcription start site (TSS). The CRISPRi system requires delivery of two components to the cell of interest: dCas9-SALL1-SDS3 protein and a CRISPRi guide RNA.
CRISPRi dCas9-SALL1-SDS3 repressor construct
In the CRISPRi system, the native Cas9 DNA-cutting functionality has been obliterated by point mutations, and the deactivated Cas9 (dCas9) protein has been further engineered for gene repression by fusing our proprietary SALL1 and SDS3 repressor domains.
We tested many repressor constructs before creating our dCas9-SALL1-SDS3 fusion. While many current CRISPRi systems rely on dCas9-KRAB variants (Gilbert et al.,2013,Yeo et al., 2018, Moghadam et al., 2020, Alerasool et al., 2020), our novel repressor, covalently linked to the dCas9 protein, further inhibits target gene transcription by recruiting proteins involved in chromatin remodeling and gene silencing (Alland et al. 2002). The result is a broadly functional repressor, turning gene expression down but not off (figure 2).
We also compared the specificity of dCas9-SALL1-SDS3 to dCas9-KRAB using whole transcriptome RNA sequencing (RNA-seq) and observed a consistent trend across the targeted genes: while both repressors were similarly highly specific, dCas9-SALL1-SDS3 mediated more potent target gene repression (figure 3). We believe our repressor system provides researchers with more distinct experimental outcomes without introducing additional noise or uncertainty.
CRISPRi guide RNA
All CRISPRmod guide RNAs are algorithm optimized to maximize specificity and performance, and are available in both synthetic and lentiviral packaged formats.
CRISPRi requires guide RNA designs that target regions 0-300 base pairs downstream of the target gene’s transcriptional start site (TSS) to result in interference. Designing functional guide RNAs can be challenging, as TSSs are not always well annotated or inaccessible due to other protein factors. Dharmacon CRISPRi guide RNA products use a published CRISPRi v2.1 algorithm developed via machine learning techniques (M. A. Horlbeck et al.,2016). The algorithm used FANTOM and Ensembl databases to predict the TSSs and incorporate chromatin, position, and sequence data to predict highly effective guide RNA designs. For challenging genes with alternative TSSs, guide RNAs that target the respective TSSs are provided. The three guide RNA designs per TSS are ranked in order of predicted functionality.
CRISPRi synthetic sgRNA
We are proud to be the first to offer CRISPRi synthetic single-guide RNA (sgRNA). This format enables co-transfection or electroporation with CRISPRi dCas9-SALL1-SDS3 mRNA or introduction into stable dCas9-SALL1-SDS3 expressing cells, and provides the fastest method for obtaining CRISPRi results. With this method, gene repression is evident 24 hours after transfection and persists through 96 hours (figure 4).
CRISPRi gene repression is observed 24 hours post-transfection and is maximal 48-72 hours post-transfection
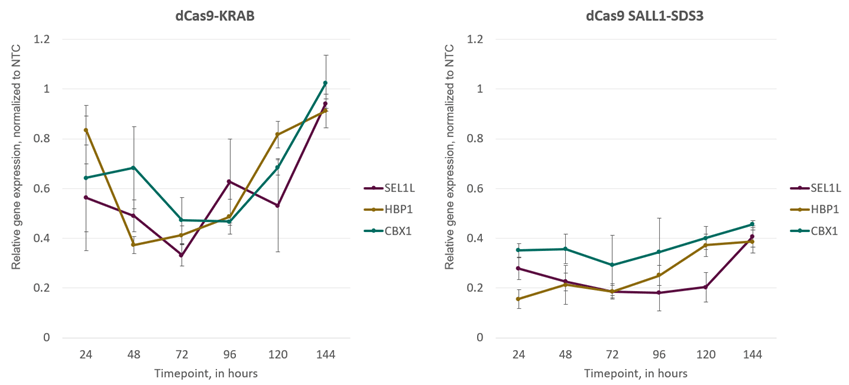
Figure 4. CRISPRi synthetic sgRNAs achieve maximal repression at 48-72 hours post-transfection in both dCas9-KRAB and dCas9-SALL1-SDS3-expressing cells. U2OS cells stably expressing integrated dCas9-KRAB or dCas9-SALL1-SDS3 were plated at 10,000 cells/well and transfected using DharmaFECT 4 Transfection Reagent with pools of pre-designed synthetic sgRNA (25nM) targeting CBX1, HBP1, or SEL1L. Cells were harvested at 24, 48, 72, 96, 120, and 144 hours post-transfection, total RNA was isolated, and relative gene expression was measured using RT-qPCR. The relative expression of each target gene was calculated with the ∆∆Cq method using GAPDH as the housekeeping gene and normalized to a non-targeting control (NTC).
Pooling synthetic sgRNA increases CRISPRi efficiency
Pooled sgRNAs produce gene knockdown equivalent or greater than the most functional individual guide RNA (figure 5).
Pooling is an excellent strategy to drive maximal gene repression and is also beneficial to decrease an experiment’s scale, for example, when performing an analysis of multiple genes in an arrayed plate format.
Pooling sgRNAs enhances gene knockdown
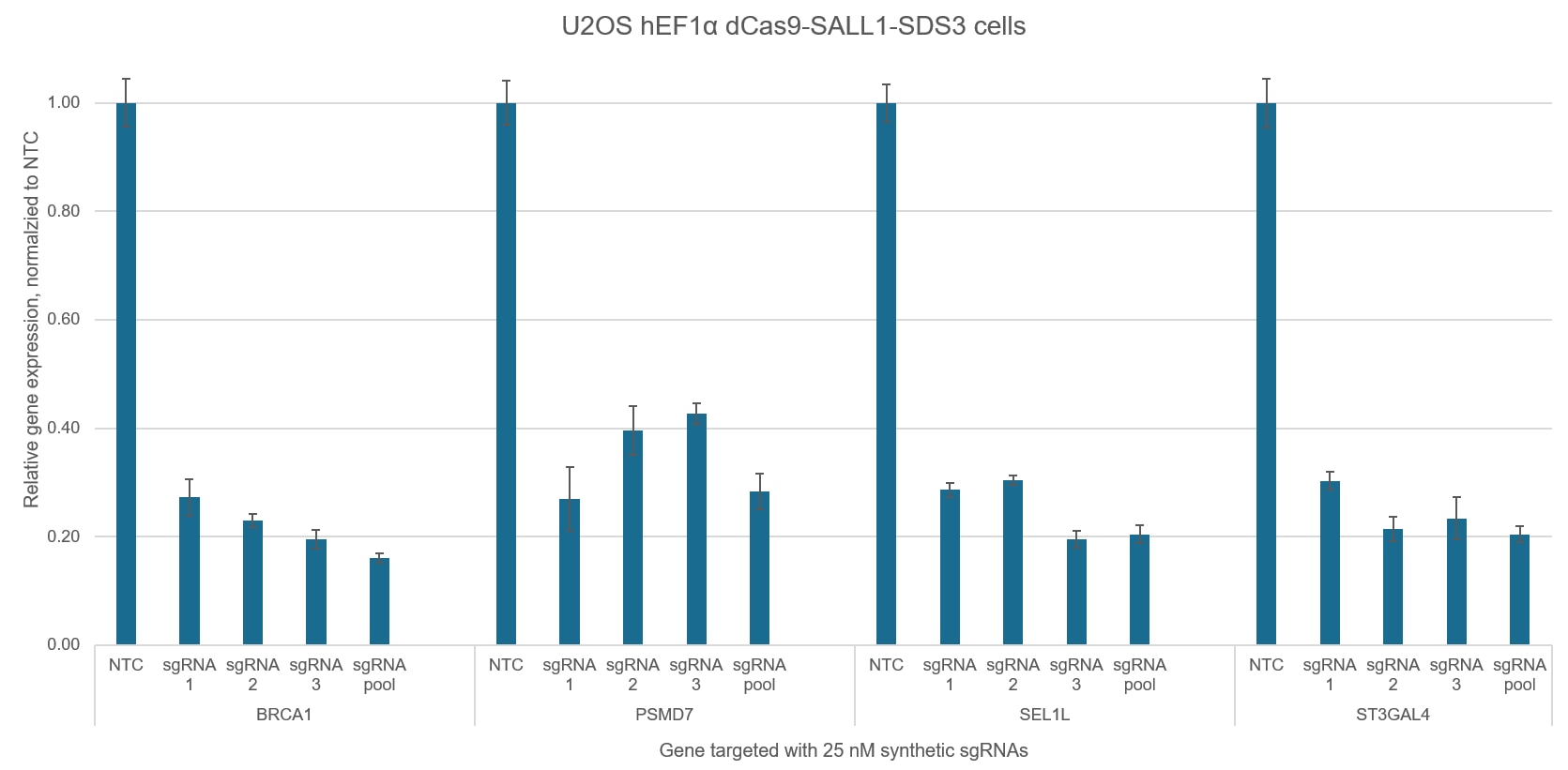
Figure 5. Individual CRISPRi sgRNAs achieve robust target gene repression independently, but when pooled together in a single reagent, display enhanced repression levels. U2OS cells stably expressing integrated dCas9-SALL1-SDS3 were plated at 10,000 cells/well and transfected using DharmaFECT 4 Transfection Reagent with CRISPRi synthetic sgRNA targeting BRCA1, PSMD7, SEL1L, or ST3GAL4. CRISPRi sgRNAs were used either individually or pooled (to a total concentration of 25 nM). Cells were harvested 72 hours post-transfection, total RNA was isolated, and relative gene expression was measured using RT-qPCR. The relative expression for each gene was calculated with the ∆∆Cq method using GAPDH as the housekeeping gene and normalized to non-targeting control (NTC).
Multiplexed gene knockdown
CRISPRi synthetic sgRNA offers a unique method for the simultaneous knockdown of multiple genes (figure 6). Each guide RNA binds to its specific DNA target independently and relies on exogenously supplied dCas9 fusion protein, thereby minimizing any competition for endogenous pathways. Our scientists have validated the effectiveness of multiplex CRISPRi in iPSC cells.
Synthetic sgRNAs enable easy multiplexing for the simultaneous repression of multiple genes
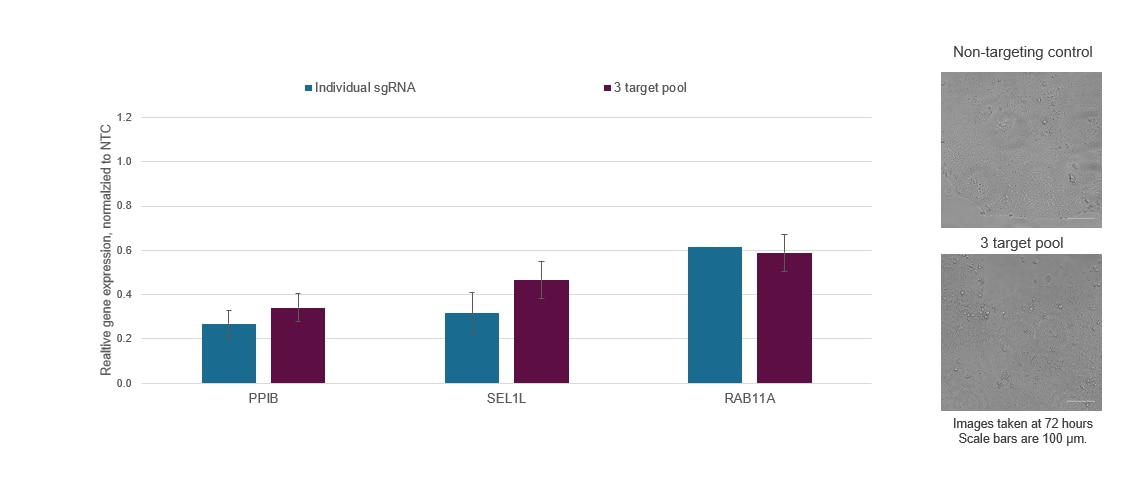
Figure 6. Individual CRISPRi sgRNAs can be pooled together in a single reagent to achieve enhanced target gene repression or multiplexed to enable the simultaneous repression of multiple genes. WTC-11 human iPS cells stably expressing integrated dCas9-SALL1-SDS3 were nucleofected with CRISPRi synthetic sgRNA targeting PPIB, SEL1L, and RAB11A via a Lonza 96-well Shuttle system. The most active CRISPRi sgRNA for each gene target was selected and used either individually or as part of a multi-target pool at a concentration of 3 µM per guide. Cells were harvested 72 hours post-nucleofection, total RNA was isolated, and relative gene expression was measured using RT-qPCR. The relative expression for each target gene was calculated with the ∆∆Cq method using ACTB as the housekeeping gene and normalized to a non-targeting control (NTC). Three genes were simultaneously repressed without a substantial decrease in target gene repression or marked changes in cell viability and morphology.
Confirming gene repression
There are many methods to confirm gene repression, including RT-qPCR, Western blots, or immunofluorescence analysis. Typically, the fastest and easiest way to measure the relative change in target gene expression levels between samples treated with a non-targeting control and CRISPRi guide RNAs is RT-qPCR. RT-qPCR analysis works with either the SYBR green method or probe-based gene expression assays. One thing to note when performing RT-qPCR is the gene expression level may drop to a level that is not detectable after CRISPRi treatment. In this case, when using the ∆∆Cq method of analysis, an arbitrary value representing the qPCR instrument’s detection limit is used as a place holder for “non-detectable” as a non-zero value is necessary to perform the calculation. In most cases, this value will be between 35 and 40, depending on the number of programmed cycles and the instrument Cq determination method. Many protocols recommend adding additional cycles (up to 45 total) to standard qPCR cycling conditions.
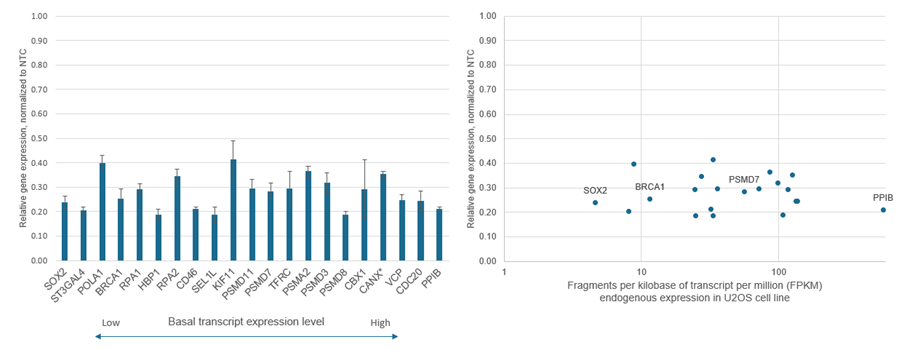
In our labs, the level of gene knockdown has been shown no correlation with the basal target expression level of each respective gene (figure 7). For example, the gene PPIB is a member of the peptidyl-prolyl cis-trans isomerase (PPIase) family. PPIB is highly expressed in many cell types and is one of the genes that we recommend as a positive control. Here we show relative gene knockdown of a wide range of genes. Interestingly, we see similar transcriptional repression across genes with high or low basal expression levels.
CRISPRi does not depend on endogenous expression levels
Figure 7. CRISPRi transcriptional repression varies by gene but does not appear to be dependent on endogenous expression levels. Data has been compiled from multiple transfections performed in U2OS cells stably expressing integrated dCas9-SALL1-SDS3 with the following matching conditions. Cells were plated at 10,000 cells/well and transfected using DharmaFECT 4 Transfection Reagent with CRISPRi synthetic sgRNA pools (25 nM) targeting 21 genes with different basal levels of expression. Cells were harvested 72 hours post-transfection, total RNA was isolated, and relative gene expression was measured using RT-qPCR. The relative expression of each target gene was calculated with the ∆∆Cq method using GAPDH as the housekeeping gene and normalized to a non-targeting control (NTC). The CRISPRi-mediated target gene repression is shown in the left graph where the genes are ordered from low to high level of basal transcript expression. In the right graph target gene expression is plotted against fragments per kilobase of transcript per million (FPKM), a representation of basal gene expression based on RNA-Seq data from the parental U2OS cell line. In U2OS cells PPIB is expressed at a basal level roughly one hundredfold greater than that of SOX2, but both genes can be repressed to roughly 20-25 percent of their basal expression with CRISPRmod CRISPRi reagents.
Orthogonal validation
Validated science is good science. Dharmacon Reagents include a wide range of reagents that interrogate cellular pathways, allowing inactivation of a gene at the DNA level, modulating RNA transcription, or by degrading mature mRNA transcripts. We are here to help you choose the best method of gene activation or inactivation for your application. Better yet, choose two methods for robust, elegant, validated science.
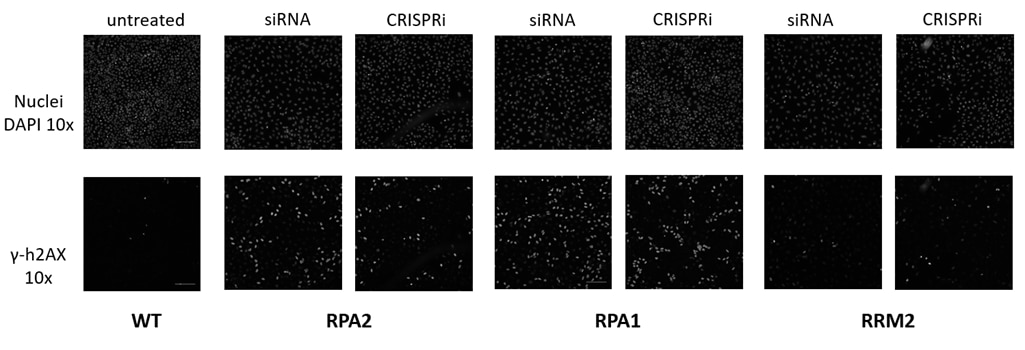
CRISPRi presents an ideal opportunity for follow-on studies after RNAi or CRISPRko screening (figure 8). Here we used CRISPRi to follow up on an arrayed siRNA screen we had previously conducted.
Synthetic CRISPRi reagents can be used to orthogonally validate hits from siRNA screens
Figure 8. Damage Response Assay. h4AX (a constituent of core histone complexes) surrounding double-strand breaks is rapidly phosphorylated, producing a set of responses through the DNA damage signaling network. Knockdown of proteins critical to the DNA damage pathway results in an accumulation of unrepaired double-stranded DNA breaks, and an increase in phosphorylated h4AX (γ-h4AX). Hits from a prior RNAi screen were orthogonally validated using CRISPRi reagents. U2OS cells constitutively expressing dCas9-SALL1-SDS3 under the hEF1α promoter were transfected with pooled sgRNAs (50nM) or ON-TARGETplus SMARTpools (50nM) targeting RPA2, RPA1, or RRM2 using DharmaFECT 4 Transfection Reagent. 72 hours post-transfection, the cells were fixed and stained with an anti-phospho-h4AX antibody, and Hoechst stain was used to identify nuclei. A duplicate plate was harvested, total RNA was isolated, and relative gene expression was measured using RT-qPCR. The relative expression for each target gene was calculated with the ∆∆Cq method using GAPDH as the housekeeping gene and normalized to non-targeting control (NTC).
Experimental considerations for CRISPRi
There are many options and considerations for CRISPRi experimental conditions. Generally, users achieve the most robust knockdown when working with a stable population of dCas9-SALL1-SDS3 cells. For extended timepoint assays (more than 120 hours), many choose lentiviral sgRNA. For short-term assays (less than 120 hours), synthetic sgRNA gives more robust gene repression than expressed guide RNAs.
| CRISPRi dCas9-SALL1-SDS3 source | Guide RNA format | Delivery method | Benefits & Recommended uses |
|---|---|---|---|
|
CRISPRi dCas9-SALL1-SDS3 lentiviral particles transduction & selection for CRISPRi dCas9 stable cells |
CRISPRi synthetic sgRNA | Transfection or electroporation |
|
| CRISPRi lentiviral sgRNA particles | Transduction |
|
|
| CRISPRi dCas9-SALL1-SDS3 mRNA | CRISPRi synthetic sgRNA | Co-transfection or electroporation |
|
CRISPRi workflows
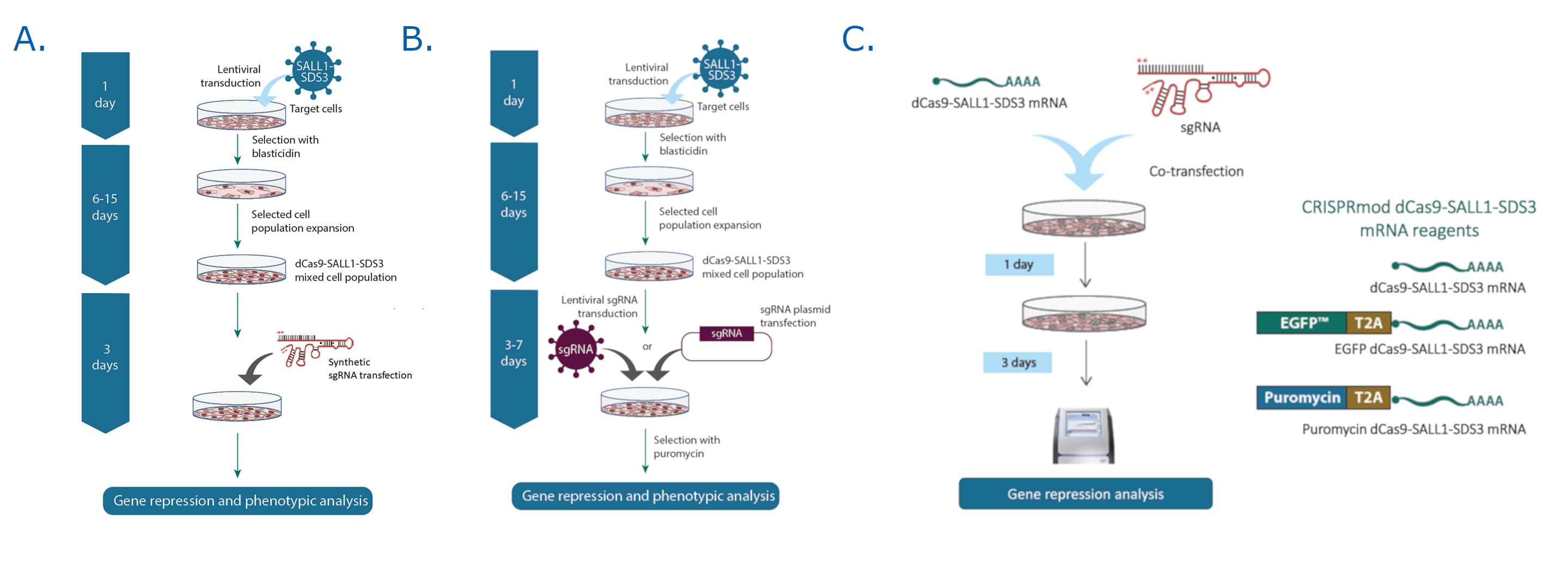
We offer multiple workflow options when using CRISPRi for transcriptional repression
- Transduce with dCas9-SALL1-SDS3 lentiviral particles to generate CRISPRi-ready, stable dCas9-SALL1-SDS3 expressing cells. Then introduce either A) synthetic or B) lentiviral sgRNA to knockdown your target gene(s). This system is ideal for screening, extended time point assays or when working with difficult-to-transfect cell types.
- C) Co-transfect or electroporate dCas9-SALL1-SDS3 mRNA with synthetic sgRNA, then enrich cell populations using fluorescence or puromycin resistance options. This system is best suited for rapid, transient gene repression studies.
References
- L.A. Gilbert, et al., CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell. 154(2): p. 442-51 (2013).
- NC. Yeo, et al., An enhanced CRISPR repressor for targeted mammalian gene regulation. Nat Methods. 15(8): p. 611-616 (2018).
- F. Moghadam, et al., Synthetic immunomodulation with a CRISPR super-repressor in vivo. Nat Cell Biol. 22(9): p. 1143-1154 (2020).
- N. Alerasool, et al., An efficient KRAB domain for CRISPRi applications in human cells. Nat Methods. 17(11): p. 1093-1096 (2020).
- L. Alland, et al., Identification of mammalian Sds3, an integral component of the Sin3? Histone deacetylase corepressor complex. Mol and cell bio. 22(8): p.2743-2750 (2002).
- M. A. Horlbeck et al., Compact and highly active next-generation libraries for CRISPR-mediated gene repression and activation. eLife. 5, e19760 (2016).
Order products
CRISPRi dCas9-SALL1-SDS3
Purified dCas9-SALL1-SDS3 mRNA or lentiviral particles offer CRISPRi solutions for any CRISPRi workflow
CRISPRi sgRNA
Synthetic and lentiviral expressed CRISPRi sgRNAs for highly specific transcriptional repression of any human gene
CRISPRmod controls
Positive and non-targeting controls to ensure your CRISPRi or CRISPRa experimental conditions
Helpful resources
CRISPRi research poster
CRISPR-mediated transcriptional repression with a novel dCas9 fusion protein and synthetic sgRNAs
Article – What is CRISPRa and CRISPRi?
CRISPR-Cas9 for gene knockdown and up-regulation