Dharmacon Edit-R CRISPR guide RNAs are guaranteed to edit the target gene-of-interest, so you can be confident in your knockout result – see guarantee tab for details.
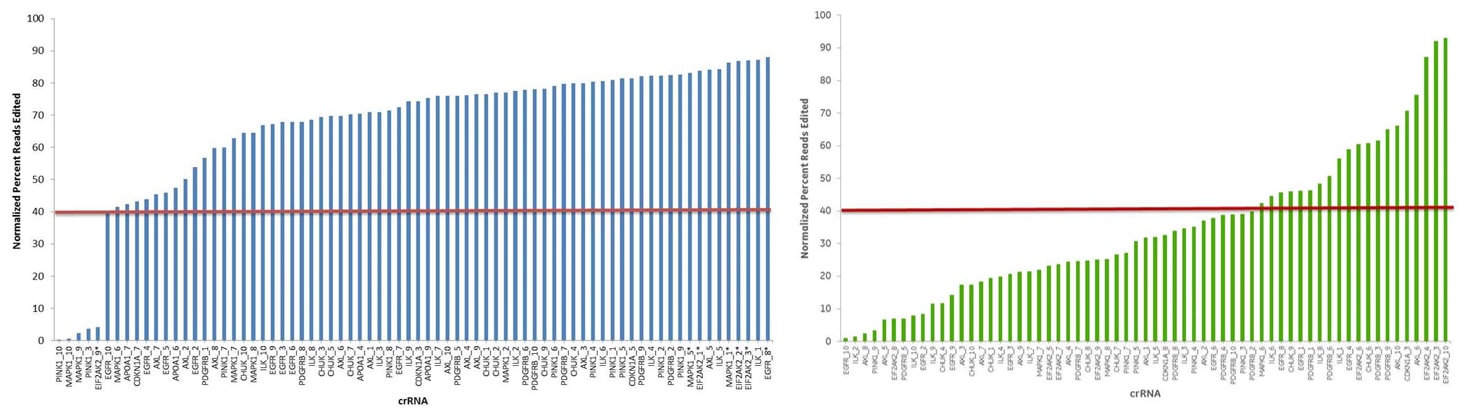
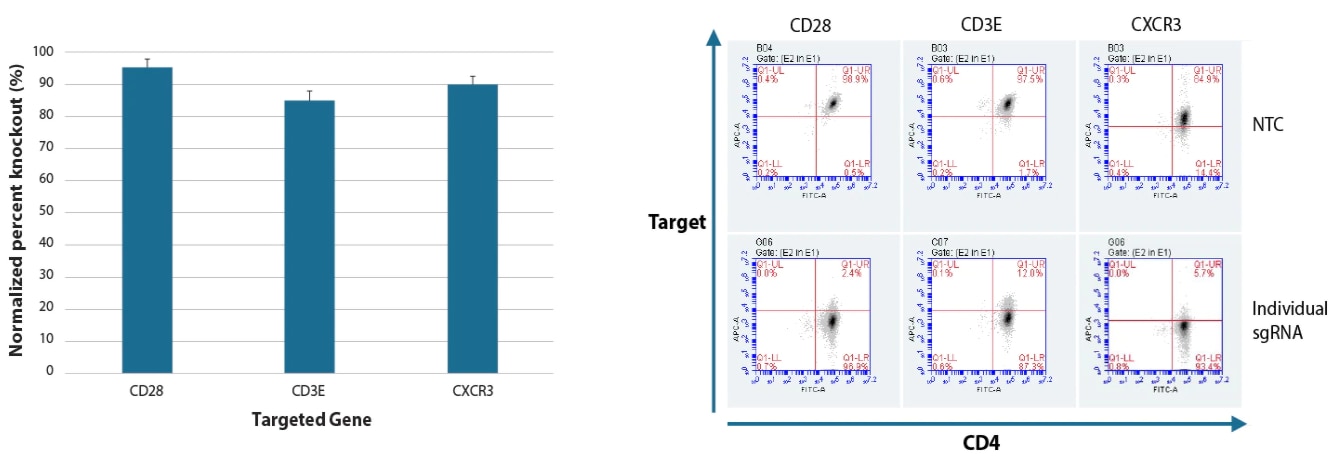
Our guide RNAs are designed using a validated algorithm to achieve functional gene knockout with high specificity. By assessing phenotypes for thousands of designs, then validating our design rules in multiple assay systems, we have established standards for determining optimal knockout target sites.
New! Edit-R human sgRNA designs have been updated to the latest RefSeq in 2025 providing the most specific and genomically relevant guides for producing efficient protein knockout. This allows the Edit-R algorithm to target the latest genome annotations more accurately and efficiently providing you with the best solution for your research needs. Please reach out to Scientific Support if you have any questions.
Visit our application page to learn more about the Edit-R CRISPR-Cas9 gene editing system.
Flexibility to fit your workflow
We offer Dharmacon Edit-R CRISPR guide RNA in pools or as individual reagents in a variety of formats to fit many different experimental workflows.
- Synthetic single guide (sgRNA) or two part crRNA:tracrRNA reagents can be co-transfected with Cas9 mRNA or protein for DNA-free workflows.
- Lentiviral sgRNA are recommended for editing in difficult-to-transfect cells.
- All-in-one lentiviral sgRNA + Cas9 offers a single-reagent knockout workflow.
- Custom design synthetic or lentiviral guide RNAs for additional species, alternative nucleases or any target location using the CRISPR Design Tool.
Scale to meet any scope
Expand the scope of your next loss-of-function experiment. Use the Cherry-Pick Library Tool to design and order your own custom CRISPR knockout screening library of Edit-R guide RNA reagents. Or see our predefined CRISPR knockout libraries for gene families or whole genome screens.
Order CRISPR guide RNA
Synthetic sgRNA
Transfection-ready synthetic sgRNAs eliminate cloning and in vitro transcription steps
Synthetic crRNA
Must be used with synthetic tracrRNA
All-in-one sgRNA + Cas9
Stable guide RNA expression for cell types not amenable to transfection
Lentiviral sgRNA
Stable guide RNA expression for cell types not amenable to transfection
Custom guide RNA solutions
CRISPR Design Tool
Design and order custom guide RNA for any species, nuclease or cut site
Custom guide RNA
When you already know your guide RNA sequence and need to get it synthesized
Guide RNA reagent selection
Determining the most appropriate guide RNA format for your experiment will depend on your particular application and cell type. Use this table to select the right reagents for your specific experimental conditions.
| Synthetic crRNA | Synthetic sgRNA | Lentiviral sgRNA | Lentiviral All-in-one sgRNA | |
|---|---|---|---|---|
| Guaranteed to edit your gene of interest | ✔ | ✔ | ✔ | ✔ |
| Can be custom designed for any species, nuclease or editing location | ✔ | ✔ | ✔* | ✔* |
| RNP compatible for electroporation | ✔ | ✔ | ||
| Transduce into difficult-to-edit cells that can't be electroporated | ✔ | ✔ | ||
| Population enrichment with antibiotic resistance markers | ✔ | ✔ | ||
| Population enrichment with fluorescent markers | ✔** | ✔** | ✔ | |
| Single reagent workflow | ✔ |
*custom vector designs for use with S. pyogenes Cas9 only.
**when co-transfected with fluorescent Cas9 mRNA.

CRISPR guide RNA
Dharmacon Edit-R synthetic guide RNA & controls
-
Synthetic sgRNA
Genome wide synthetic sgRNA designs guaranteed to edit your gene of interest. The design algorithm maximizes the potential for functional protein knockout while minimizing off-target editing through stringent specificity checks.
-
Synthetic sgRNA positive controls and detection primers
Species-specific synthetic sgRNAs targeting well-characterized genes, as well as mismatch detection assay primers, to determine the effectiveness of your gene editing conditions for maximal efficiency.
-
Synthetic sgRNA non-targeting controls
Non-targeting controls to evaluate cellular responses to CRISPR-Cas9 components in the absence of gene target-specific sgRNA.
-
Cutting controls
Cutting control (safe harbor) synthetic sgRNAs recommended for determination of baseline cellular responses in CRISPR-Cas9 experiments.
-
T-cell receptor targets
Validated Edit-R crRNAs targeting T cell receptor genes: TRAC, TRDC, and TRBC.
-
-
Synthetic crRNA
Guaranteed to edit your target! Algorithm-optimized crRNA for genome-wide coverage of human and mouse genes. Modifications for nuclease resistance improve DNA-free editing.
-
Positive controls and detection primers
Species-specific crRNAs targeting well-characterized genes, as well as mismatch detection assay primers, to determine the effectiveness of your gene editing conditions for maximal efficiency.
-
Non-targeting controls
Non-targeting controls to evaluate cellular responses to CRISPR-Cas9 components in the absence of gene target-specific crRNA.
-
Cutting controls
Cutting control (safe harbor) crRNAs recommended for determination of baseline cellular responses in CRISPR-Cas9 experiments.
-
T-cell receptor targets
Validated Edit-R crRNAs targeting T cell receptor genes: TRAC, TRDC, and TRBC.
-
Synthetic tracrRNA
Trans-activating CRISPR RNA (tracrRNA) is required for use with Edit-R synthetic crRNA to form the complex that programs Cas9 nuclease.
Dharmacon Edit-R lentiviral sgRNA & controls
-
All-in-one lentiviral sgRNA + Cas9
Combined Cas9 and sgRNA expression for efficient gene knockout & unparalleled specificity; available as glycerol stocks and high-titer purified particles.
-
All-in-one lentiviral sgRNA positive controls and kits
All-in-one lentiviral sgRNA controls to verify DNA double-strand breaks and gene editing efficiencies.
-
All-in-one lentiviral sgRNA non-targeting controls
All-in-one lentiviral sgRNA constructs bioinformatically designed to not target any gene in human or mouse genomes.
-
Cutting Controls
Cutting control (safe harbor) sgRNA recommended for determination of baseline cellular responses in CRISPR-Cas9 experiments.
-
Lentiviral sgRNA
Guaranteed to edit your target, algorithm-optimized sgRNA for genome-wide coverage of human or mouse genes. Provided as high-titer lentiviral particles or glycerol stocks.
-
Lentiviral sgRNA positive controls
Species-specific sgRNAs targeting well-characterized genes to determine the effectiveness of your gene editing conditions for maximum efficiency.
-
Lentiviral sgRNA non-targeting controls
Non-targeting controls to evaluate cellular responses to CRISPR-Cas9 components in the absence of gene-specific sgRNA.
-
Cutting Controls
Cutting control (safe harbor) sgRNA recommended for determination of baseline cellular responses in CRISPR-Cas9 experiments.
Custom guide RNA design
-
CRISPR Design Tool
Place a custom guide RNA order, or design and order your own synthetic sgRNA, crRNA, or lentiviral sgRNA with our easy-to-use interface.
Dharmacon Edit-R HDR donor template design, ordering tools & kits
-
HDR Donor Designer - oligo
Design and order a single-strand DNA donor (≤ 150 nt) for insertion, deletion, or other alteration.
-
HDR Donor Designer - plasmid
Design and order a plasmid DNA donor kit for insertion of a mKate2 or EGFP fluorescent marker, or other a custom insert.
-
HDR plasmid donor kits
Rapidly and easily assemble a plasmid donor for HDR
-
HDR plasmid donor components
Quickly and efficiently build a HDR donor plasmid
-
HDR plasmid donor primers
PCR components for Edit-R Plasmid Donor Kits
Cas9 and dCas9 nuclease sources
"CRISPR-ready" premade Cas9-expressing cell lines
-
Cas9 expressing stable cell lines
Choose from a variety of popular cell backgrounds, then simply deliver a Edit-R CRISPR guide RNA targeted to any gene for simple loss-of-function studies.
Transient Cas9 nuclease for DNA-free workflows
-
Cas9 nuclease mRNA
Purified Cas9 mRNA for transient Cas9 nuclease expression. Fluorescent options available for sorting, enrichment and visualization of delivery.
-
Cas9 nuclease protein NLS
Purified Cas9 protein ready-to-use for DNA-free workflows.
Vector-based solutions for generating Cas9-expressing cell lines
-
Lentiviral Cas9 nuclease reagents
Purified lentiviral particles or plasmid DNA for the generation of stable Cas9 nuclease-expressing cell populations. Constitutive or inducible promoter options are available. -
Strict-R inducible Cas9 lentiviral system
Lentiviral Cas9 system for dual-controlled gene knockout in diverse cell types.
CRISPR knockout screening libraries
-
Custom cherry-pick libraries
Have a favorite gene list? Customize and order plates of synthetic or lentiviral guide RNAs for knockout studies in your targets of interest.
-
CRISPR libraries
Pre-defined collections of synthetic and lentiviral guides covering popular gene families or whole genomes in pooled and arrayed format.
The Dharmacon Edit-R guarantee
We guarantee that EVERY predesigned guide RNA will provide successful editing at the target site when delivered as described in the Edit-R technical manuals.
The Edit-R guide RNA guarantee is valid when used with any wild type S. pyogenes Cas9 nuclease, including mRNA, expression plasmid, protein, or stable Cas9 expression, and Edit-R crRNAs must be used with Edit-R tracrRNA for the guarantee to apply.
Analysis of editing of the treated cell population must be shown using a T7EI or Surveyor mismatch detection assay. If successful editing is not observed for a predesigned Edit-R guide RNA while an appropriate side-by-side Edit-R positive control is successful, a one-time replacement of a different predesigned Edit-R guide RNA of the same format and quantity will be provided at no cost.
A replacement will only be approved upon discussion with our Scientific Support team.
Successful editing at the DNA level does not always lead to functional gene knockout; it is recommended to test multiple guide RNAs to determine the most effective guide RNA for knockout of your target gene.
This guarantee does not extend to any accompanying experimental costs, does not apply to guide RNAs ordered via the CRISPR Design Tool, and will not be extended to the replacement guide RNA.

