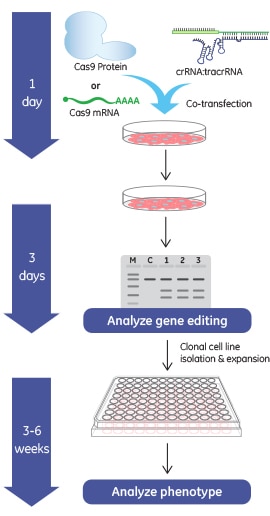
What is CRISPR-Cas technology?
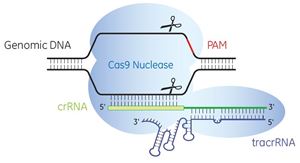
Short RNAs can be used to guide nuclease proteins to targets within complex eukaryotic genomes with high specificity, based upon a naturally occurring system for bacterial and archaeal immunity. Popularly known as CRISPR-Cas (Clustered Regularly Interspersed Short Palindromic Repeats) technology, these systems rely upon CRISPR-associated (Cas) proteins and have the potential to carry out genome editing functions or alter gene expression. The components of CRISPR-Cas9 genome editing systems can be combined in multiple ways for various gene editing applications.
What applications are possible with CRISPR-Cas systems?
Gene knockout
The Cas9 endonuclease has become a popular tool for directed gene editing in eukaryotic systems1-3. With the use of a target-specific CRISPR RNA (crRNA) and trans-activating crRNA (tracrRNA), or a fused format called a single guide RNA (sgRNA), locations within complex mammalian genomes can be targeted by the Cas9 endonuclease for a double stranded break1. These breaks can be repaired by endogenous DNA repair mechanisms through a process known collectively as non-homologous end-joining (NHEJ). Because NHEJ is error prone, genomic deletions or insertions (indels) can result that create frame shifts and premature termination to permanently silence target genes. It is important to be aware that the insertions and deletions resulting from NHEJ are random and differ from cell to cell. The exact genomic changes that result can be determined by additional experiments on clonal cell lines. The crRNA, tracrRNA, and sgRNAs can either be transcribed intracellularly, in vitro transcribed or custom synthesized and introduced through transfection. Intracellular expression of Cas9 endonuclease can be accomplished by plasmid or integrated lentiviral expression vectors driven by constitutive or inducible promoters. Effective levels of intracellular Cas9 can also be delivered by DNA-free systems that have the advantage of exceedingly low possibility of additional genomic alterations, through transfection of mRNA encoding Cas9 or Cas9 protein delivery.
DNA-free CRISPR-Cas9 gene editing
What does “DNA-free” CRISPR-Cas9 gene editing really mean? It means that your system uses no CRISPR-Cas9 components in the form of DNA vectors; each component is either RNA or protein. Starting with Cas9 mRNA or purified Cas9 protein as the source for Cas9 nuclease expression in genome engineering experiments has advantages for some applications. Why? The use of DNA-based Cas9 or guide RNA expression systems carries with it the possibility of undesirable genetic alterations due to plasmid DNA integration at the cut site or random lentiviral vector integrations. For this reason, a DNA-free gene editing system can be a good choice for creating engineered cell lines. If your experiment involves observation of a phenotype in a non-clonal cell population, you may not need a DNA-free option. However, if your experimental end-goal does not require enrichment of Cas9-expressing cells and you wish to avoid potential integration events, then consider using Cas9 mRNA or purified Cas9 protein.
Homology-directed repair (HDR)
The CRISPR-Cas9 induced double-strand break can also be used as an opportunity to create a knockin, rather than a target gene knockout. The precise insertion of a donor template can alter the coding region of a gene to “fix” a mutation, introduce a protein tag, or create a new restriction site. We have demonstrated that single-strand DNA can be used to create precise insertions using synthetic crRNA and tracrRNA with Cas9 nuclease. Alternatively, the activity of Cas9 can be altered to nick rather than perform a double stranded cut. The Cas9 nickase can be used with a pair of crRNA:tracrRNA complexes or sgRNAs targeted to two closely spaced regions on opposite strands, and when used with a short double stranded DNA, homology-directed repair can performed2,4.
- Read more details on HDR in this Application Note Homology-directed repair with Dharmacon™ Edit-R™ CRISPR-Cas9 reagents and single-strand DNA oligos.
- Download a scientific poster on the use of ssDNA and synthetic guide RNAs for HDR
Transient gene silencing or transcriptional repression (CRISPRi)
With this application, Cas9 is modified so that it cannot cut DNA, and when combined with a guide RNA that targets a promoter region, the complex can reduce transcriptional activity and concomitant gene expression5,6. Read more about CRISPRi here.
Transient activation of endogenous genes (CRISPRa or CRISPRon)
By employing a Cas9 mutant that cannot cut DNA and to which a transcriptional activation domain has been fused, the expression of endogenous genes can be up-regulated by targeting the Cas9 fusion protein to the promoter region of an endogenous target gene, or multiple genes simultaneously6,7. Read more about CRISPRa here.
Embryonic stem cell and transgenic animals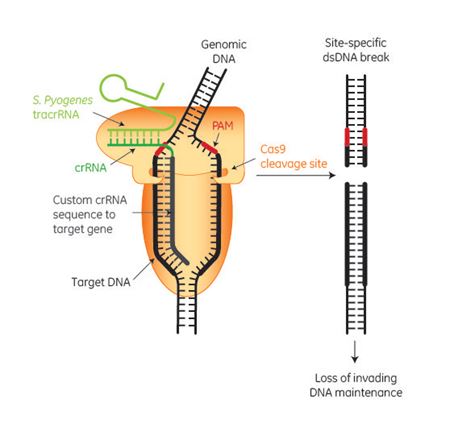
CRISPR-Cas systems can be used to rapidly and efficiently engineer one or multiple genetic changes to murine embryonic stem cells for the generation of genetically modified mice8. A similar approach has been used to genetically modify primate single cell embryos9.
Pooled genome-scale knockout screening
Pooled lentiviral libraries have been used at low MOI to perform genome-scale screens of cancer cell viability, pluripotency, and drug resistance10,11. With single copy number integration, bi-allelic gene silencing can be accomplished with high frequency. Relative representation of genes involved in producing the phenotypic criteria of the screen (e.g. cell death, proliferation, drug resistance, etc.) can be identified by next generation sequencing.
- Read this brief overview on performing functional genomics screens using CRISPR-Cas9 lentiviral sgRNA pooled screening.
- Download this protocol for pooled screening with lentiviral Cas9 and sgRNAs.
The importance of CRISPR-Cas9 controls for experimental design
Positive controls
For almost every CRISPR-Cas9 gene editing application, the use of one or more positive controls is an essential first step. Transfection efficiencies vary between cell types, and must be optimized for each to achieve high-efficiency gene editing. Guide RNAs that are known to edit with high efficiency and/or target a sensitive region of a gene for a clear phenotypic readout should be used as positive controls. The outcome of the positive control experiment should be evaluated by a mismatch detection assay to confirm presence of indels, and quantify editing efficiency for each of the transfection conditions evaluated. Once optimal conditions have been established, experimental gene editing can begin using the same high-efficiency conditions determined by the positive control.
Negative controls
Negative controls are especially important for CRISPR-Cas9 experiments designed to be evaluated by phenotypic readouts. Changes in cell phenotype, viability or gene expression levels in cells treated with negative controls likely reflect a baseline cellular response that can be compared to the levels in cells treated with target-specific crRNAs for more accurate interpretation of results.
Synthetic sgRNA for CRISPR-Cas9 experiments
Although a natural synthetic two RNA (crRNA:tracrRNA) system is very efficient and cost-effective for most applications, researchers working with in vivo and ex vivo models have indicated a preference for a sgRNA system.
- Learn more about CRISPR-Cas9 genome editing with synthetic 99-mer single guide RNA
CRISPR-Cas9 guide RNA algorithm
To identify criteria for choosing the best functional guide RNA, we developed an algorithm based on functional gene knockout using a phenotypic assay for proteasome function with a GFP readout. Get an overview of how the Edit-R algorithm was developed to select guide RNAs more likely to cause functional protein knockout.
CRISPR-Cas9 guide RNA specificity
Identification of potential off-target cleavage sites is essential for CRISPR specificity. Learn more about the alignment tools and strategies available.
Inducible lentiviral Cas9 nuclease
The timing of any gene editing experiment can be critical, especially when setting up a pooled screen, generating a stable cell line without risk of increased metabolic load, or integrating the Cas9 in a precursor state and performing an experiment in a later derived cell state. An inducible Lentiviral Cas9 Nuclease can provide the temporal control necessary to ensure editing only occurs when it is required.
References:
- Jinek, M., et al., RNA-programmed genome editing in human cells. Elife, 2013. 2: p. e00471.
- Cong, L., et al., Multiplex genome engineering using CRISPR/Cas systems. Science, 2013. 339(6121): p. 819-23.
- Mali, P., et al., RNA-guided human genome engineering via Cas9. Science, 2013. 339(6121): p. 823-6.
- Ran, F.A., et al., Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell, 2013. 154(6): p. 1380-9.
- Qi, L.S., et al., Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell, 2013. 152(5): p. 1173-83.
- Gilbert, L.A., et al., CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell, 2013. 154(2): p. 442-51.
- Cheng, A.W., et al., Multiplexed activation of endogenous genes by CRISPR-on, an RNA-guided transcriptional activator system. Cell Res, 2013. 23(10): p. 1163-71.
- Wang, H., et al., One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell, 2013. 153(4): p. 910-8.
- Niu, Y., et al., Generation of Gene-Modified Cynomolgus Monkey via Cas9/RNA-Mediated Gene Targeting in One-Cell Embryos. Cell, 2014. 156(4): p. 836-43.
- Wang, T., et al., Genetic screens in human cells using the CRISPR-Cas9 system. Science, 2014. 343(6166): p. 80-4.
- Shalem, O., et al., Genome-scale CRISPR-Cas9 knockout scre
Order products
Cas9 nuclease
Configure the optimal promoter for your cell type to ensure robust Cas9 expression or explore DNA-free options
CRISPR controls and detection primers
Proper controls are essential to assessment of CRISPR-Cas9 genomic editing experiments
CRISPR guide RNA
High quality, ready-to-use lentiviral and synthetic reagents to guide Cas9 cleavage
CRISPR-Cas9 gene editing
Optimized tools for high-confidence genome engineering
CRISPR-Cas9 screening libraries
Pooled sgRNA or arrayed crRNA for high-throughput gene editing studies
Helpful resources
A CRISPR-Cas9 gene engineering workflow - App Note
The CRISPR-Cas9 system is being widely used for genome engineering in many different biological applications
Edit-R CRISPR-Cas9 Gene engineering with Cas9 nuclease expression plasmids and synthetic RNAs - Tech Manual
Dharmacon™ Edit-R™ CRISPR-Cas9 gene engineering with Cas9 nuclease expression plasmids and synthetic guide RNAs
Video - Edit-R CRISPR-Cas9 gene engineering platform
Learn how to choose the right products for your CRISPR gene knockout workflow, cell type(s) and experimental goals. This video walks through a few high-level questions to help direct you to the best components for your experiment.
The CRISPR-Cas9 system is being widely used for genome engineering in many different biological applications.