RNA synthesis
In a cell, RNA is made by the transcription of DNA by intracellular enzymes. RNA can also be made in vitro by enzymes like T7 RNA polymerase. A third option for making RNA exists - chemical synthesis - which differs from in vivo and in vitro RNA production because it does not use enzymes to make RNA. Instead, chemical synthesis works by adding one RNA base at a time in repeated cycles. In this way, short single-strand RNA molecules, or RNA oligonucleotides, of exactly defined sequence and length are made
2'-ACE explained
Since 1995 Dharmacon has been a leader of RNA synthesis technology through the development of a novel RNA synthesis chemistry that is so advanced we patented it! This unique chemistry uses a new class of silyl ethers to protect the 5'-hydroxyl with a unique acid-labile orthoester protecting group on the 2'-hydroxyl [1, 2]. The chemical name of this protecting group is 2´-bis(acetoxyethoxy)-methyl ether, but we commonly refer to it as “2'-ACE”. To learn more, read our 2'-ACE RNA Synthesis Chemistry Technical Note which explains our patented RNA synthesis chemistry in greater detail.
2'-ACE advantages
The innovative properties of 2´-ACE technology enable routine synthesis of siRNA in high yield and of exceptional quality. This is because 2´-ACE phosphoramidites allow high coupling efficiencies resulting in high quality RNAs and siRNAs. The purity levels from this synthesis technology, typically 80-85% for unmodified siRNA, are routinely higher than other methods (e.g. 2'-tert-Butyldimethylsilyl ether, TBDMS). Other advantages of 2'-ACE protecting groups are water solubility, nuclease resistance, synthesis of strong secondary structures, and a final acid deprotection that is mild, fast, and requires minimal handling.
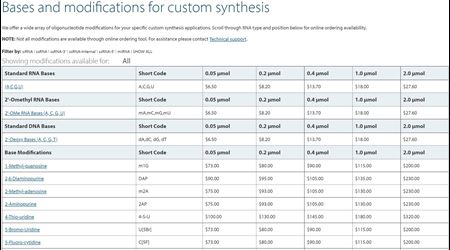
See all available custom options and pricing details »
What are the benefits of chemical modification for siRNA?
Chemical synthesis of siRNAs allows for position-specific incorporation of a wide variety of modifications into your sequence. These modifications can provide siRNA performance advantages for various experimental applications [3, 4].
Stability - resistance to nuclease degradation
The simplest approach to increase nuclease resistance of siRNA is to directly modify the internucleotide phosphate linkage. Replacement of a non-bridging oxygen with sulfur (phosphorothioate, PS) has been extensively used to improve nuclease resistance of siRNAs [5].
Modifications of the 2'-position of the ribose can indirectly improve nuclease resistance of the internucleotide phosphate bond while increasing duplex stability (Tm). This modification has also been shown to provide protection from immune activation.
A combination of moderate PS backbone modifications with small, well-tolerated 2'-substitutions (2'-O-Methyl, 2'-Fluoro, 2'-Hydro) has often created highly stable siRNAs for applications in vivo [6].
Potency - transient silencing duration
The transient nature of siRNA silencing is primarily due to the continuous dilution of intracellular siRNA through cellular division and nucleic acid degradation. Chemical modifications help to ensure delivery of more intact siRNAs into the cell, and subsequently stabilize the delivered siRNA for proper RISC recruitment. For example, prolonged silencing has been achieved in cell culture using 2'-OMe- modifications within the siRNA [7].
Immunogenicity - induction of innate immune responses
One major challenge for in vivo siRNA is undesired activation of the innate immune system. The immunostimulatory effects of dsRNA are mediated primarily through three Toll-like receptors (TLR3, TLR7, and TLR8) and by proteins such as retinoic acid inducible protein (RIG-1) [8], oligoadenylate synthetase (OAS), dsRNA-responsive kinase (PKR), and melanoma differentiation associated protein (MDA-5) [9].
Rendering siRNAs unrecognizable to these immunostimulatory receptors through chemical modifications has been shown to significantly decrease siRNA immunogenicity. Modifying specific sequences with 2'-OMe, 2'-F, and 2'-H can effectively reduce TLR7/TLR8 interaction while generally preserving silencing activity [10, 11]. Additional modifications, such as 2-thiouracil, pseudouracil, 5-methylcytosine, 5-methyluracil, and N6-methyladenosine have also been shown to minimize the immune effects mediated by TLR3, TLR7, and TLR8 [12].
Specificity - preventing off-target effects
When either strand of an siRNA reduces the expression of a gene that is not the intended target - this is undesirable and known as “off-targeting”. There are two ways off-targeting can occur. The siRNA guide strand can target an mRNA transcript from a gene that has a sequence similar to the target gene. A second way off-targeting occurs is through loading of the siRNA passenger strand into RISC, and degradation of mRNA targets that have sequences complementary to the passenger strand. Both of these off-targeting mechanisms usually occur through high degrees of sequence similarity in the seed region, and imperfect matches throughout the rest of the siRNA oligonucleotide. For this reason, these off-targeting events are considered to be microRNA-like in nature [13].
To prevent either siRNA strand from miRNA-like off-targeting, position-specific modifications are used.
- Modifications in the seed region of the siRNA guide strand are used to reduce off-targeting potential.
- Introduction of chemical modifications in duplexes that result in low seed Tm values potentially eliminate off-target effects with minimal negative impact on siRNA efficiency [14].
- Chemical modifications may also be employed to prevent loading of the passenger strand into RISC by altering the thermodynamic asymmetry of the duplex so as to bias for guide strand loading. For example, a single unlocked nucleic acid (UNA) modification at the 5'-terminus of the passenger strand can reduce its incorporation into RISC without hindering siRNA guide strand loading [15].
- A number of modified nucleotide bases (5-bromo, 5-iodo, 2-thio, 4-thio, dihydro-, and pseudo-uracil) have also been used to stabilize base-pairing and enhance base-pairing specificity [16, 17].
Delivery - cellular uptake
Chemical modification can help solve some of the problems associated with delivery of siRNA into certain cells and animals. Due to the negatively charged phosphate backbones of RNA and its analogues, and the non-polar nature of the lipid bi-layer that makes up the cell membrane, getting siRNA into a cell can be difficult. For systemic delivery of siRNA, various conjugates have been shown to help delivery of siRNA into cells. These modifications include:
- cholesterol
- tocopherol and folic acid
- linkers
- lipids
- polymers
- peptides
- aptamers
These modifications can increase the potential use of siRNA as a systemic drug [18].
Imaging - probes for monitoring experimental performance
For some siRNA experiments, imaging siRNA can be a useful control for delivery, or to determine the pharmacokinetics (PK) and pharmacodynamics (PD) of the molecules in the experimental model.
Here are multiple ways imaging probes can be used:
- To determine what tissues take up RNA molecules introduced in vivo, 18F or 64Cu isotopes can be conjugated to siRNAs and used for positron emission tomography (PET) imaging [19, 20].
- Single-proton emission computed tomography (SPECT) using 111In- or 99mTc-labeled siRNA have been applied to define the PK and biodistribution of siRNAs [21, 22].
- Bioluminescence imaging, commonly used to quantify luciferase expression from reporter systems, can provide a reliable assay system for quantifying the functionality of siRNAs in vivo or for accessing the PD [23].
- Aside from bioluminescence, in vivo fluorescent imaging can detect the presence of near-infrared (NIR) dyes through mammalian cells. Fluorescence has been used to study the biodistribution patterns of Cy5- or Cy5.5-labeled siRNAs [24, 25].
Summary
The best modification to use in your custom siRNA design depends upon your intended application and can vary with the design of siRNA, specific sequence, and method of delivery. Chemical modifications in RNA synthesis can widen the experimental approaches of RNAi that can be taken. The vast array of chemical modification schemes offered by Dharmacon will greatly extend your research capabilities. If you have questions about chemical modifications for custom RNA synthesis, speak with a scientist on our Scientific Support Team.
References
- Scaringe, S.A. et al. (1998) Novel RNA synthesis method using 5´-silyl-2´-orthoester protecting groups. J. Am. Chem. Soc. 120:11820-11821.
- Scaringe, S.A. et al. (2004) Preparation of 5´-silyl-2´-orthoester ribonucleosides for Use in oligoribonucleotide synthesis. Curr. Protoc. Nuc. Acid Chem. 2.10.1-2.10.16.
- Rana T.M. (2007) Illuminating the silence: understanding the structure and function of small RNAs. Nature Reviews Mol. Cell Bio. 8:23-36.
- Bramsen, J.B. and Kjems J. (2012) Development of therapeutic-grade small-interfering RNAs by chemical engineering. Frontiers in Genetics 3:1-22.
- Kole R. (2012) RNA therapeutics: beyond RNA interference and antisense oligonucleotides. Nature Reviews Drug Discovery 11:125-140.
- Soutschek, J. et al. (2004) Therapeutic silencing of an endogenous gene by systemic administration of modified siRNAs. Nature 432:173-178.
- Volkov, A.A. (2009) Selective protection of nuclease-sensitive sites in siRNA prolongs silencing effect. Oligonucleotides 19:191-202.
- Hornung, V. et al. (2006) 5’-Triphosphate RNA is the ligand for RIG-1. Science 314:994-997.
- Robbins, M. et al. (2009) siRNA and innate immunity. Oligonucleotide 19:89-101.
- Judge A.D. et al. (2006) Design of noninflammatory synthetic siRNA mediating potent gene silencing in vivo. Mol. Ther. 13:494-505.
- Cekaite, L. et al. (2007) Gene expression analysis in blood cells in response to unmodified and 2’-modified siRNAs reveals TLD-dependent and independent effects. J. Mol. Biol. 365:90-108.
- Kariko, K. et al. (2005) Suppression of RNA recognition by toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23:165-175.
- Birmingham, A. et al. (2006) 3’ UTR seed matches, but not overall identity, are associated with RNAi off-targets. Nature Methods 3:199-204.
- Ui-Tei, K. et al. (2008) Thermodynamic stability and Watson-Crick base pairing in the seed duplex are major determinants of the efficiency of the siRNA-based off-target effect. Nucleic Acids Res. 36:7100-7109.
- Snead, N.M. et al. (2013) 5’ Unlocked nucleic acid modification improves siRNA targeting. Mol. Ther. Nucleic Acids 2:e103.
- Chiu, Y.L. and Rana, T.M. (2003) siRNA function in RNAi: a chemical modification analysis. RNA 9:1034-1048.
- Sipa, K. et al. (2007) Effect of base modifications on structure, thermodynamic stability, and gene silencing activity of short interfering RNA. RNA 13:1301-1316.
- Winkler J. (2013) Oligonucleotide conjugates for therapeutic applications. Ther. Deliv. 4:791-809.
- Viel, T. et al. (2008) Molecular imaging study on in vivo distribution and pharmacokinetics of modified small interfering RNAs (siRNAs). Oligonucleotides 18:201-212.
- Bartlett, D.W. et al. (2007) Impact of tumor-specific targeting on the biodistribution and efficacy of siRNA nanoparticles measured by multimodality in vivo imaging. Proc. Natl. Acad. Sci. USA 104:15549-15554.
- Malek, A. et al. (2009) In vivo pharmacokinetics, tissue distribution and underlying mechanisms of various PEI(-PEG)/siRNA complexes. Toxicol. Appl. Pharmacol. 236:97-108.
- Liu, N. et al. (2007) Radiolabeling small RNA with technetium-99m for visualizing cellular delivery and mouse distribution. Nucl. Med. Biol. 34:399-404.
- Bartlett D.W. and Davis M.E. (2006) Insights into kinetics of siRNA-mediated gene silencing from live-cell and live animal bioluminescent imaging. Nucleic Acids Res. 34:322-333.
- Huang, Y. et al. (2011) Elimination pathways of systemically delivered siRNA. Mol. Ther. 19:381-385.
- Whitehead, K.A. et al. (2014) Degradable lipid nanoparticles with predictable in vivo siRNA delivery activity. Nature Commun. 5:4277.
Order products
Custom DNA Synthesis
Order a wide range of single-strand DNA to support your HDR donor oligo needs or other experimental constructs.
Custom microRNA Modulation Tools
Modify an existing microRNA mimic or inhibitor, target a novel microRNA, or request an Accell microRNA Inhibitor or Stabilized Mimic for special applications.
Custom siRNA Synthesis
siRNA customization has never been easier
Services
Use our online design tools and extensive synthesis options to create custom products specific to your application.
siDESIGN Center
Design and order siRNAs targeting genes in non-standard species, particular splice variants, or homologous regions across gene families or species.
Single-strand RNA Synthesis
Create and order RNA molecules with a wide variety of chemical modifications or learn about our capabilities for long RNA oligos, dye labeling and custom amidites.
Helpful resources
2'-ACE RNA Synthesis Chemistry
A new class of silyl ethers is used to protect the 5’-hydroxyl (5’-SIL) in combination with an acid-labile orthoester protecting group
Chemical Synthesis of Long and Highly Modified RNA
A significant improvement in RNA synthesis technology, 2'-ACE chemistry results in higher yields, greater purity and superior ease of handling.
Custom RNA - Resources
Find product guides, FAQs and more.
In vivo RNAi: Biodistribution, Delivery, and Applications
The dsRNA is cleaved by the RNase-III enzyme, Dicer, into small interfering RNAs (siRNA) that are approximately 21-23 nt.