CRISPR activation (CRISPRa) allows researchers to up-regulate specific gene function by activating transcription, without editing the DNA.
Visit our CRISPRa applications page to learn more.
The Dharmacon™ CRISPRmod CRISPRa system requires two components to operate: a CRISPRa guide RNA and a catalytically deactivated Cas9 (dCas9) fused to transcriptional activators (VPR). Our CRISPRa reagents provide a straightforward, efficient set of tools to study a gene’s function by overexpression in its native context.
CRISPRa reagent selection
There are many options and considerations for CRISPRa experimental conditions. Generally, users achieve the most robust activation when working with a stable population of dCas9-VPR expressing cells.
| dCas9-VPR source | Guide RNA format | Delivery method | Recommendations for use & benefits |
|---|---|---|---|
|
CRISPRa dCas9-VPR stable cells or CRISPRa dCas9-VPR lentiviral particle transduction |
CRISPRa synthetic sgRNA | Transfection or electroporation |
|
| CRISPRa synthetic crRNA + tracrRNA | |||
| CRISPRa lentiviral sgRNA particles | Transduction |
|
|
| CRISPRa dCas9-VPR mRNA | CRISPRa synthetic sgRNA | Co-transfection or electroporation |
|
| CRISPRa synthetic crRNA + tracrRNA | |||
| CRISPRa dCas9-VPR lentiviral plasmid | CRISPRa lentiviral sgRNA plasmid | Co-transfection or electroporation with dCas9-VPR plasmid |
|
| CRISPRa all-in-one dCas9-VPR + sgRNA | Transduction |
|
|
| Strict-R inducible lentiviral dCas9-VPR CRISPRa | Transduction |
|
|
As you explore CRISPR-mediated gene activation, consider whether the NEXTFLEX™ Rapid Directional RNA Kit 2.0 could complement your gene modulation analysis through additional transcriptome profiling.
Order CRISPRa guide RNA
CRISPRa lentiviral sgRNA
An ideal delivery method for CRISPR activation in difficult-to-transfect cells.
CRISPRa synthetic sgRNA
Predesigned synthetic guide RNA for over-expression of human and mouse genes using CRISPR activation.
CRISPRa synthetic crRNA
Co-transfect with tracrRNA for a DNA-free CRISPR activation workflow.
CRISPRa All-in-one Lentiviral sgRNA
Single reagent – dCas9-VPR + sgRNA in one vector for easy gene to cell delivery.
CRISPRa dCas9 solutions
dCas9-VPR stable cell lines
"CRISPRa-ready" cell lines - simply introduce guide RNA for targeted gene activation.
dCas9-VPR mRNA
Option to co-express fluorescent marker or antibiotic resistance for straightforward enrichment.
dCas9-VPR lentiviral reagents
Lentiviral dCas9-VPR vectors for robust gene activation in a broad range of relevant cell types.
Strict-R inducible lentiviral CRISPRa system
Lentiviral dCas9-VPR system for dual controlled gene activation in diverse cell types
SAM transcriptional activation system
MS2 tracrRNA
For use with synthetic CRISPRa crRNA and a stable cell line expressing both components (NLS-dCas9-VP64 and MS2-p65-HSF1) of the SAM activation system.
Helpful Resources
Watch our "cDNA vs. CRISPRa" video
Which is better for your overexpression experiment? We’ll give some tips to help you choose.
Watch our "What is inducible vs constitutive gene expression?" video
Gene expression using plasmid vectors is either inducible or constitutive. Explore what this means using products from our CRISPRa portfolio as examples.
CRISPRa synthetic guide RNA & controls
-
CRISPRa synthetic sgRNA
Offering coverage of the human and mouse genomes, these synthetic RNAs are modified for nuclease resistance and available as pooled or individual sgRNA reagents.
-
CRISPRa synthetic sgRNA positive controls
Pooled or individual sgRNA controls for assessment of optimal experimental conditions for gene activation
-
CRISPRa synthetic sgRNA non-targeting controls
Non-targeting controls to evaluate baseline cellular responses to CRISPRa components in the absence of gene target-specific sgRNA
-
CRISPRa synthetic crRNA
Offering coverage of the human and mouse genomes, these synthetic RNAs are modified for nuclease resistance and available as pooled or individual crRNA reagents.
-
CRISPRa synthetic crRNA positive controls
Pooled or individual crRNA controls for assessment of optimal experimental conditions for gene activation
-
CRISPRa synthetic crRNA non-targeting controls
Non-targeting controls to evaluate baseline cellular responses to CRISPRa components in the absence of gene target-specific crRNA
-
Synthetic tracrRNA
Trans-activating CRISPR RNA (tracrRNA) is a synthetic, HPLC-purified, long RNA required for use with crRNA to form the complex that programs dCas-VPR. It is modified for nuclease resistance.
:
CRISPRa lentiviral guide RNA & controls
-
CRISPRa lentiviral sgRNA
Lentiviral particles or glycerol stocks of vector-based single guide RNA are predesigned for genome-wide coverage of human and mouse genes
-
CRISPRa lentiviral sgRNA positive controls
Control sgRNAs targeting well-characterized genes to determine the effectiveness of your experimental conditions for maximum activation
-
CRISPRa lentiviral sgRNA non-targeting controls
Non-targeting controls to evaluate baseline cellular responses to CRISPRa components in the absence of target-specific sgRNA
CRISPRa dCas9-VPR reagents
-
dCas9-VPR expressing stable cell lines
"CRISPRa ready" cells in a variety of commonly used backgrounds - simply introduce CRISPRa guide RNA for targeted gene activation.
-
dCas9-VPR mRNA
Co-transfect with CRISPRa synthetic guide RNA for a lentiviral-free workflow.
-
dCas9-VPR lentiviral particles
Used to generate dCas9-VPR expressing stable cell lines.
CRISPRa screening libraries
-
Custom cherry-pick libraries
Upload your own gene list to customize and order plates of predesigned crRNA for CRISPRa studies across tens or thousands of genes.
-
CRISPRa synthetic sgRNA libraries
Arrayed collections of synthetic CRISPR RNA for overexpression screening of gene families, druggable, and whole human genome..
-
CRISPRa All-in-one sgRNA library
Pooled CRISPRa all-in-one lentiviral human whole genome library
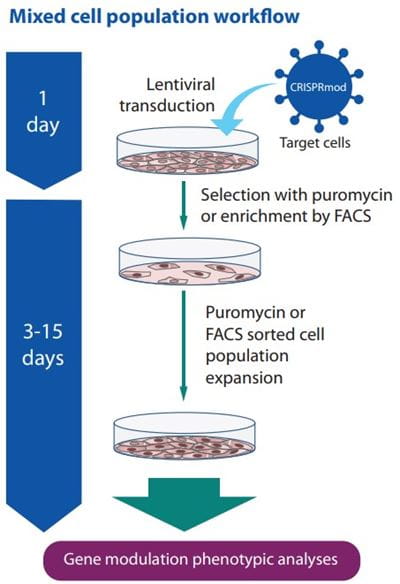
CRISPRa workflow diagram with stable dCas9-VPR expression

CRISPR activation workflow with Lentiviral dCas9-VPR and synthetic crRNA:tracrRNA (right side) or Lentiviral expressed sgRNA (left side).