Yeast Protein Interactome Collection
The Yeast Protein Interactome Collection allows for genome-wide in vivo screening of protein-protein interactions (PPI) in Saccharomyces cerevisiae.


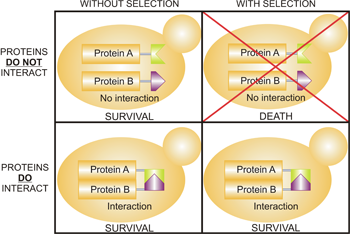
First published in Science, Tarassov et al. used the Yeast Protein Interactome Collection to interrogate protein networks in Saccharomyces cerevisiae for systems-level behavior. This collection's high sensitivity is based on protein-fragment complementation and can detect as little as 25 to 100 complexes per cell. Using colony growth as an indication of PPI, this simple, but elegant selection strategy allowed the authors to systematically test over 15 million matings. The Yeast Protein Interactome Collection, with its published interactome, provides a basis to further explore individual networks or to test for genetic and environmental effects on a genome scale.
We provide certain clone resources developed by leading academic laboratories. Many of these resources address the needs of specialized research communities not served by other commercial entities. In order to provide these as a public resource, we depend on the contributing academic laboratories for quality control.
Therefore, these are distributed in the format provided by the contributing institution "as is" with no additional product validation or guarantee. We are not responsible for any errors or performance issues. Additional information can be found in the product manual as well as in associated published articles (if available). Alternatively, the source academic institution can be contacted directly for troubleshooting.
The stock plates will be provided in 96-well microtiter plates. These will ship on dry-ice via overnight delivery and should be stored at –80°C immediately upon receipt. Please contact our customer service department for a quote and estimated shipping time on bulk orders.