- Gene editing
- Pin-point™ Reagents
- Pin-point™ Synthetic sgRNA Validated Control
Pin-point™ Synthetic sgRNA Validated Controls
Synthetic sgRNA controls to verify optimal parameters for precise gene editing without inducing DNA double-strand breaks.
Each control sgRNA is chemically synthesized, HPLC purified, and biologically validated for optimal use with either Adenine base editor (ABE) or Cytosine base editor (CBE) Pin-point base editing mRNAs.

|
Pin-point synthetic sgRNA are 128-nucleotide chimera fusions of the crRNA, tracrRNA, and a proprietary aptameric sequence. They are chemically modified for nuclease resistance on both 5’ and 3’ ends of the molecule. The sgRNA controls are recommended as positive controls for base editing experiments utilizing Pin-point ABE or Pin-point CBE mRNAs. Gene-specific positive controls and kits are designed and validated for assessment of base editing and phenotypic protein knockout where applicable. They are configured as follows:
|
Design and order custom CBE or ABE Pin-point sgRNA now
|
Base editing with the Pin-point platform induces specific nucleotide changes without the formation of DNA double-strand breaks or indels.
This system consists of three components:
[1] a nuclease-defective "nickase" nCas9 that only cuts or “nicks” a single strand of DNA. For cytosine base editing (CBE), the nCas9 is fused to a sequence of uracil glycosylase (UGI) inhibitors (Komor, 2016). For adenine base editing (ABE), the nCas9 is not fused to any UGI.
[2] a cytidine deaminase (rAPOBEC) or an adenine deaminase (ABE-flex or ABE-exact) fused to an aptamer binding protein. The rAPOBEC enzyme converts C-G base pairs to T-A base pairs if the base pairs are within the editing window, positions 4 to 8 at the PAM-distal end of the protospacer (Collantes, 2021). ABE-flex converts A-T base pairs to G-C base pairs if the base pairs are within positions 4 to 8 of the base editing window while ABE-exact converts A-T base pairs to G-C base pairs if the base pairs are within positions 5 to 6 of the base editing window.
[3] an aptameric single guide RNA (sgRNA) that recruits the nCas9 and the aptamer-deaminase fusion to a specific DNA target site.
Delivery of all three components into mammalian cells induces the base editing conversion at the targeted site. This system can be used to make gene knockouts through the introduction of premature stop codons (Billon, 2017), the disruption of splice donor and splice acceptor sites (Webber, 2019), or to introduce point mutations. See our Pin-point base editing sgRNA design tool to design your own sgRNAs for gene knockout.
|
|
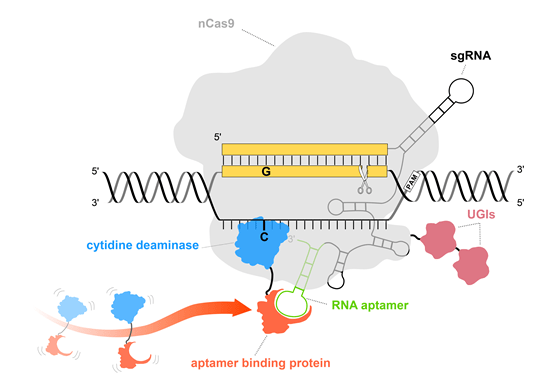
Illustration of Pin-point base editing system.
The utilization of nCas9 (light gray) with or without UGIs (red) ensures that no DNA double-strand breaks occur and DNA damage response pathway is not triggered. Pin-point sgRNA contains an aptamer (green) that is used to recruit deaminase (blue = cytidine deaminase, turquoise = adenine deaminase) via aptamer binding protein (orange) to perform base editing.
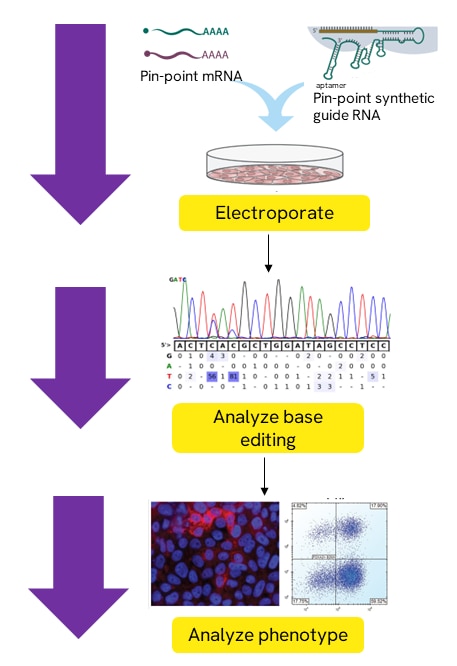
Base editing with Pin-point ABE or CBE nCas9 mRNA, Pin-point ABE or CBE deaminase mRNA, and Pin-point synthetic sgRNA
All three components are electroporated into a specific cell type of interest. Base editing may then be observed using analysis primers to amplify and sequence the base editing region of interest. A phenotype may be analyzed to assess target protein knockout where applicable.
Table 1: How much Pin-point sgRNA do I need?
| Number of reactions by well/format size* | ||||||||
| T cells (250,000 cells per well in a 24 well plate) |
iPSCs (100,000 cells per well in a 12 well plate) |
HSPCs (50,000 cells per well in a 96 well plate) |
HEK 293T cells (50,000 cells per well in a 96 well plate) |
|||||
| CBE |
ABE | CBE | ABE | CBE | ABE | CBE | ||
| 20 pmol/well |
20 pmol/well
|
40 pmol/well | 40 pmol/well | 62.5 pmol/well | 62.5 pmol/well | 60 pmol/well | ||
| 2 nmol | 100 | 100 | 50 | 50 | 32 | 32 | 33 | |
| 5 nmol | 250 | 250 | 125 | 125 | 80 | 80 | 83 | |
| 10 nmol | 500 | 500 | 250 | 250 | 160 | 160 | 167 | |
Table 2: How much Pin-point ABE mRNA do I need?
|
Number of reactions by well/format size*
|
||||||||||
| T cells (250,000 cells per well in a 24 well plate) |
iPSCs (100,000 cells per well in a 12 well plate) |
HSPCs (50,000 cells per well in a 96 well plate) |
||||||||
| ABE nCas9 mRNA | ABE-exact deaminase mRNA | ABE-flex deaminase mRNA | ABE nCas9 mRNA | ABE-exact deaminase mRNA | ABE-flex deaminase mRNA | ABE nCas9 mRNA | ABE-exact deaminase mRNA | ABE-flex deaminase mRNA |
||
| 1.5 μg/well | 1 μg/well | 0.1 μg/well | 1 μg/well | 1 μg/well | 0.1 μg/well | 1.5 μg/well |
1 μg/well | 0.4 μg/well | ||
| 10 μg (2 μg/μL) | - | 10 | 100 | - | 10 | 100 | - | 10 | 25 | |
| 20 μg (2 μg/μL) | 13 | 20 | 200 | 20 | 20 | 200 | 13 | 20 | 50 | |
| 100 μg (2 μg/μL) | 67 | 100 | 1000 | 100 | 100 | 1000 | 67 | 100 | 250 |
|
| 500 μg (2 μg/μL) |
333 | - | 500 | - | - | 333 | - | - |
||
Table 3: How much Pin-point CBE mRNA do I need?
| Number of reactions by well/format size* | ||||||||
| T cells (250,000 cells per well in a 24 well plate) |
iPSCs (100,000 cells per well in a 12 well plate) |
HSPCs (50,000 cells per well in a 96 well plate) |
HEK 293T cells (50,000 cells per well in a 96 well plate) |
|||||
| CBE nCas9 mRNA |
CBE rAPOBEC deaminase mRNA | CBE nCas9 mRNA | CBE rAPOBEC deaminase mRNA | CBE nCas9 mRNA | CBE rAPOBEC deaminase mRNA | CBE nCas9 mRNA | CBE rAPOBEC deaminase mRNA | |
| 1.56 μg/well |
0.22 μg/well
|
2.56 μg/well | 0.4 μg/well | 2.8 μg/well | 2 μg/well | 1 μg/well | 0.1 μg/well | |
| 20 μg (2 μg/μL) |
12 | 90 | 8 | 50 | 7 | 10 | 20 | 200 |
| 100 μg (2 μg/μL) | 64 | 450 | 39 | 250 | 36 | 50 | 100 | 1000 |
| 500 μg (2 μg/μL) |
320 | 2252 | 195 | 1250 | 179 | 250 | 500 | 5000 |
For additional Supporting Data please see our Pin-point Base Editing mRNA, Custom Pin-point Synthetic sgRNA and Pin-point Synthetic sgRNA Non-targeting Controls pages.

(A) Pin-point CBE nCas9 mRNA, CBE rAPOBEC deaminase mRNA, and synthetic sgRNA CBE validated controls were delivered to activated human T cells, induced pluripotent stem cells (iPSCs) and hematopoietic stem and progenitor cells (HSPCs) via electroporation. (B) Pin-point ABE nCas9 mRNA, ABE-exact deaminase mRNA, and synthetic sgRNA ABE validated controls were delivered to T cells, iPSCs and HSPCs via electroporation. (C) Pin-point ABE nCas9 mRNA, ABE-flex deaminase mRNA, and synthetic sgRNA ABE validated controls were delivered to T cells, iPSCs and HSPCs via electroporation. Base editing efficiencies were measured at 3 days post electroporation by Sanger sequencing the edited loci using Pin-point control PCR primers. N=2-4 technical replicates. Bars – average ± SD.

Figure 2: Efficient multiplex base editing with the Pin-point ABE platform in human T cells.
Activated human T cells were electroporated with Pin-point ABE nCas9 mRNA, ABE-flex deaminase mRNA, and synthetic sgRNA ABE validated controls targeting CD52, TRAC and PDCD1. (A) Base editing levels were assessed by Sanger sequencing of PCR products using Pin-point analysis primers at 3 days post electroporation. (B) Percent protein knockout was evaluated by flow cytometry at 5-6 days post electroporation. To induce the expression of PD1, T cells were cultured in the presence of phorbol12- myristate 13-acetate (PMA) and ionomycin for 48 h prior. Expression of CD52 and TCRa/b were analyzed on non-stimulated cells. Percentage of negative cells for each marker in the edited and un-edited (negative control, neg ctrl) reported as percentage of live cells. N=2 technical replicates. Bars – average ± SD.

Figure 3: Efficient multiplex base editing with the Pin-point CBE platform in human T cells.
Activated human T cells were electroporated with Pin-point CBE mRNAs and Pin-point synthetic sgRNA CBE validated controls for the 3 targets (CD52, PDCD1 and TRAC), or non-targeting control (NTC #1) sgRNA. Mock-electroporated (EP) cells were used as negative controls. (A) Cells were harvested 6-7 days post electroporation, and base editing levels were calculated from Sanger sequencing data of PCR products using the Pin-point analysis primers. N = 8 T cell donors, 5 independent experiments with duplicate or triplicate technical replicates. Bars are averages ± SD. (B) Cells were analyzed by flow cytometry 7 days post electroporation. To induce the expression of PD1, T cells were cultured in the presence of phorbol 12- myristate 13-acetate and ionomycin for 48 h prior. Expression of CD52 and TCRa/b were analyzed on non-stimulated cells. Percentage of negative cells for each marker in the edited and mock-electroporated (EP) reported as percentage of live cells. N = 2 T cell donors with duplicate or triplicate technical replicates. Bars are averages ± SD.

Figure 4: Triple knockout (KO) with the Pin-point CBE platform.
(A) Percentage of individual cells that are negative for all three markers simultaneously (CD52, PD1 and TCRa/b) in the samples in Figure 3 (edited with the Pin-point CBE platform, and mock-electroporated (EP) controls), reported as percentage of live cells, with no additional enrichment. To induce the expression of PD1, T cells were cultured in the presence of phorbol 12- myristate 13-acetate and ionomycin for 48 h prior to analysis to enable simultaneous quantification of all three markers. N = 2 T cell donors with duplicate or triplicate technical replicates. Bars are averages ± SD. (B) Representative flow cytometry plots from an edited sample to show the gating strategy: CD52 negative population gated on live cells; TCRa/b negative population gated on CD52 negative population; and PD1 negative population gated on CD52 and TCRa/b double negative population. The count of cells from this gate are the total number of cells negative for the three markers. The counts of triple negative cells are normalized on the counts of live cells and the percentage reported in panel A.
Posters
-
Performance and modularity of Revvity's Pin-point base editing system characterized by arrayed and pooled screening platforms - poster
-
Pin-point™ base editing system: a versatile editing platform driving cell therapies - poster
-
The Pin-point™ base editing platform streamlines the generation of hypoimmunogenic iPSCs for allogeneic cell therapy - poster
Safety data sheets
Selection guides
Short protocols
-
Pin-point™ CBE base editing in HEK293T and T cells, Neon Electroporation – Short Protocol
-
Pin-point™ ABE base editing in HPSCs – Short Protocol
-
Pin-point™ ABE base editing in iPSCs – Short Protocol
-
Pin-point™ ABE base editing in T cells – Short Protocol
-
Pin-point™ CBE base editing in HEK293T and T cells, Lonza 4D-Nucleofection – Short Protocol
-
Pin-point™ CBE base editing in HPSCs – Short Protocol
-
Pin-point™ CBE base editing in iPSCs – Short Protocol
Related Products
Revvity’s Pin-point™ base editing design tool makes it simple to create base editing sgRNAs that deliver reliable, high-efficiency gene knockouts.
Non-targeting synthetic controls for evaluation of Pin-Point base editing. Bioinformatically designed to not target any gene in the human genome
The Pin-point base editing mRNA system allows for precisely directed point mutation edits without inducing double stranded breaks or the need for homology directed repair

