- Gene modulation
- RNA interference
- Lincode GAS5 Control Pool
Lincode GAS5 Control Pool

Lincode GAS5 Control Pool is an effective positive control for RNAi experiments for knockdown of human long, noncoding RNAs (lncRNAs). Growth arrest-specific 5 (GAS5) is expressed during growth arrest phase of the cell cycle. The four component Lincode GAS5 siRNAs are designed to target various regions of the GAS5 transcript, providing more efficient overall knockdown results. Each siRNA carries a proprietary dual-strand modification patterns to have fewer off-targets than unmodified siRNAs. Endogenous GAS5 expression levels should be determined prior to experiments with this reagent.
Highlights
- Positive control for an endogenous lncRNA in human cells
- Chemically matched for use with Lincode siRNA reagents
- Effective silencing verified in multiple cell types
Application considerations
Prior to beginning lncRNA knockdown experiments, the expression level of GAS5, or other target lncRNA, should be assessed in your cell line. If GAS5 levels are not reliably detectable, it is recommended to use ON-TARGETplus GAPD siRNA Control Pool or ON-TARGETplus Cyclophilin B Control Pool as a means of assessing transfection and RNAi efficiency.
Sense strand activity is prevented by siRNA modifications
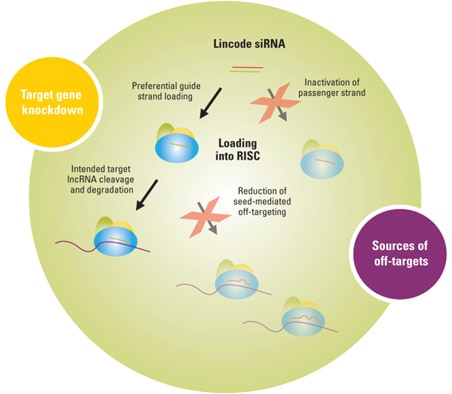
Lincode siRNAs are modified with a proprietary dual-strand modification that improves siRNA functionality. Additionally, off-targets are reduced due to:
- Inactivation of passenger strand activity; driving preferential loading of the guide strand into RISC
- Novel seed region modifications for disruption of microRNA-like off-targets
Effective silencing of GAS5 by Lincode Control siRNA Reagents
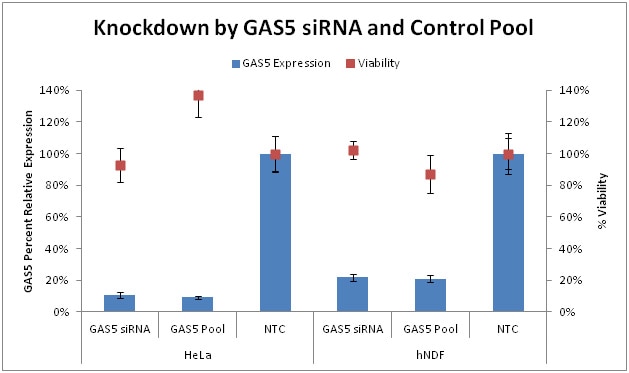
Knockdown of endogenous GAS5 in HeLa cells and hNDF cells by GAS5 Control siRNA and GAS5 Control Pool. All siRNAs were used at 25 nM. Remaining lncRNA was detected by corresponding Solaris qPCR lncRNA expression assay. Knockdown was normalized to Non-targeting control siRNA. Viability was measured by resazurin assay and normalized to Untreated cells.
Product inserts
Safety data sheets
Selection guides
Related Products
The most broadly applicable DharmaFECT formulation for optimal siRNA or microRNA transfection into a wide range of cell types for successful RNAi experiments
Catalog ID:T-2001-01
Unit Size:0.2 mL
