- Dharmacon Screening libraries
- shMIMIC Lentiviral microRNA Pooled Libraries
shMIMIC Lentiviral microRNA Pooled Libraries
Perform functional screening of hundreds or thousands of microRNAs without high-throughput automation

Optimal microRNA expression for high-confidence functional analysis screening
Target every human and mouse microRNA detailed in the miRBase database. Stable expression of a mature microRNA, using shMIMIC Lentiviral microRNA pooled libraries.
Screen large numbers of microRNAs without automation. shMIMIC pooled libraries available for all human and mouse miRBase matures, as well as a human/mouse conserved collection.
Library construction, pooling techniques, and high-throughput sequencing-compatible screening workflows validated to ensure reproducibility and accurate hit identification. Learn about optimal library design and the pooled screening workflow on our Pooled Lentiviral Screening Libraries page.
Highlights:
- Innovative vector design for robust microRNAs expression from shMIMIC lentiviral microRNAs
- SMARTchoice options - select the promoter and reporter (or no reporter!) for optimal performance in your experiment
You may also select shMIMIC Inducible microRNA pooled libraries for tight regulation of expression from the Tet-On® 3G transactivator protein
Each shMIMIC lentiviral library includes:
- 30 Non-targeting controls
- 80 species-specific positive control shRNAs targeting 10 common viability and reference genes (8 shRNAs each)
- Data file including construct sequences, miRBase gene IDs, and counts per millions of mapped reads
| shMIMIC microRNA Pooled Library | Number of mature microRNAs | Number of unique designs | Number of pools × number of constructs per pool1 |
|---|---|---|---|
| Human | 2580 | 2555 | 1 pool × 2665 constructs |
| Mouse | 1913 | 1896 | 1 pool × 2006 constructs |
| Human-Mouse Conserved | 355 human + 341 mouse | 386 | 1 pool x 496 constructs |
Reagents recommended in our validated protocol:
- Vector-matched SMARTvector Non-targeting control for transduction optimization
- SMARTvector Indexing PCR and Sequencing Primer Kits (A & B) with 12 unique indexing primers, optimized and experimentally validated for:
- Efficient PCR amplification of genomic DNA with minimal bias
- High-throughput multiplexed Next Gen Sequencing for hit identification
Before ordering, download these valuable tools to plan your pooled lentiviral screen and calculate the amounts of components required:
- SMARTvector & shMIMIC Lentiviral Pooled Libraries Technical Manual
- Pooled Lentiviral Screening Laboratory Protocols & Calculation Tracking Worksheet
Important Notice
The shMIMIC Lentiviral microRNA Pooled Libraries are solely for internal research use (as set forth in the Product Terms and Conditions) in laboratories where the containment measures stated below and in applicable laws and regulations are met. Reagents may not be used for diagnostic, therapeutic or other commercial purposes and may not to be administered to humans for any purpose or to animals for therapeutic purposes. SMARTvector Lentiviral shRNA Pooled Libraries are provided as lentiviral particles are replication-incompetent, self-inactivating (SIN) and non-pathogenic (do not cause infectious human disease).
Any investigator who purchases lentiviral particle products is responsible for consulting with their institution's health and biosafety personnel for specific guidelines on the handling of lentiviral vector particles. Furthermore, each investigator is fully responsible for obtaining the required permissions for research using and the acceptance of replication-incompetent SIN lentiviral vectors and replication-defective lentiviral particles into their local jurisdiction and institution.
Elements of the shMIMIC microRNA Lentiviral Backbone
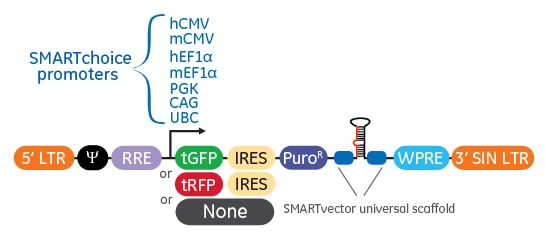
| Vector Element | Utility |
|---|---|
| 5' LTR | 5' Long Terminal Repeat necessary for lentiviral particle production and integration of the construct into the host cell genome |
| Ψ | Psi packaging sequence allows viral genome packaging using lentiviral packaging systems |
| RRE | Rev Response Element enhances titer by increasing packaging efficiency of full-length viral genomes |
| tGFP or tRFP | TurboGFP or TurboRFP reporter for visual tracking expression upon doxycycline induction |
| None | No-reporter option for use in applications where fluorescence is not required or desired |
| IRES | Internal Ribosomal Entry Site allows expression of fluorescent marker and puromycin resistance in a single transcript |
| PuroR | Puromycin resistance permits antibioticselective pressure and propagation of stable integrants |
| SMARTvector universal scaffold | Optimized proprietary scaffold in which mature microRNA sequence is embedded |
| WPRE | Woodchuck Hepatitis Post-transcriptional Regulatory Element enhances transgene expression in target cells |
| 3' SIN LTR | 3' Self-inactivating Long Terminal Repeat for increased Lentiviral safety |
shMIMIC Pooled Screening Workflow
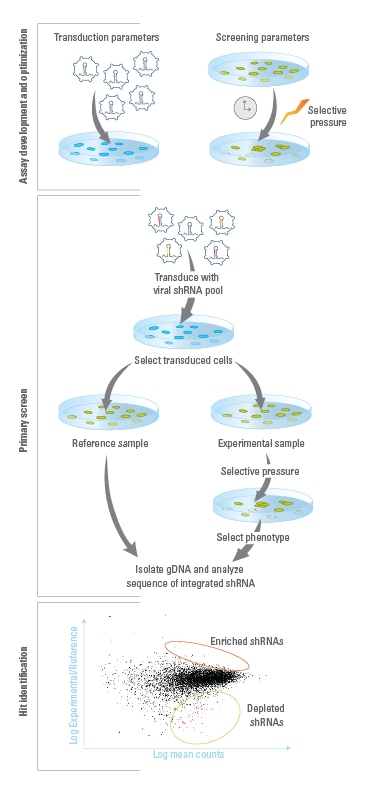
Assay Development and Optimization: Establish optimal experimental conditions, including those for a) lentiviral transduction and b) screening parameters, such as selective pressure and time between collection of reference and experimental samples.Primary Screen: A stable population of cells expressing single integrands of constructs are created by transducing lentiviral pools at low MOIs. Transduced cells are then split into reference and experimental populations for application of a selective pressure that induces the phenotype of interest. Genomic DNA (gDNA) is then isolated from reference and experimental populations of transduced cells. Illumina-adapted primers and Phusion Hot-Start II High Fidelity DNA Polymerase are used to PCR amplify integrated construct sequences and add Illumina flow-cell binding sequences. The resulting amplicons are run on Illumina platform sequencers, using the sequencing primers provided.Hit Identification and Follow-up: Construct sequences are identified in reference and experimental libraries. Constructs that are enriched or depleted during the screen are identified as hits, and the genes that they target are identified. Hits can be confirmed and studied further using individual constructs that can be ordered from the Dharmacon catalog collection.
Strength of promoter activity varies from one biological context to another and can affect experimental outcomes.
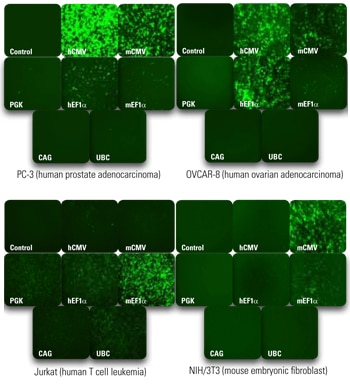
Promoter activity varies across several human and rodent cell lines. Cells were plated at a density of 50,000 cells per well in a 24-well plate and transduced at MOI = 15 with SMARTvector Empty Vector Control Particles expressing TurboGFP. Promoter activity was assessed at 72 hours post-transduction by the fluorescence intensity of TurboGFP.
The Human-Mouse Conserved microRNA Collection
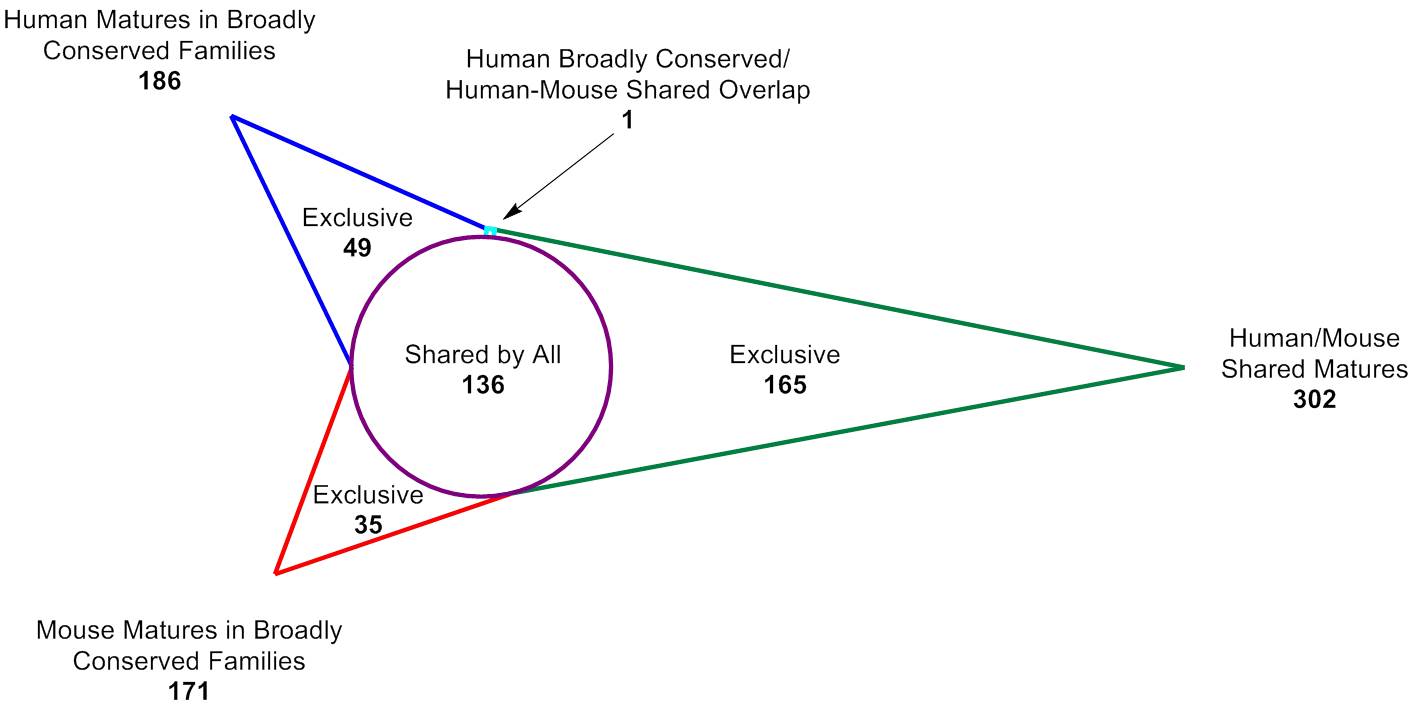
The Human-Mouse Conserved microRNA pooled library is useful for researchers that need ultra-high fold coverage for subtle phenotypes, or who may not have enough cells (e.g. primary cells) for a larger screen and wish to focus on highly conserved microRNAs. It can be used in either human or mouse models, and translates well to in vivo applications.It includes 386 total mature microRNAs:302 human and mouse matures that are identical35 mouse matures in broadly conserved families that are not identical to human49 human matures in broadly conserved families that are not identical to mouseBroadly conserved families as defined at www.targetscan.org
Strezoska, A. Licon, Optimized PCR Conditions and Increased shRNA Fold Representation Improve Reproducibility of Pooled shRNA Screens. PLoS One 7, e42341 (2012).
