- Gene modulation
- RNA interference
- Mimic Housekeeping Positive Control #2 (GAPD)
miRIDIAN microRNA Mimic Housekeeping Positive Control #2 (GAPD)
Validated miRIDAN microRNA Mimics designed to target GAPD in human, mouse and rat

miRIDIAN microRNA Mimic Positive Controls are ideal for optimizing conditions for relevant, well-controlled microRNA modulation experiments. Both positive and negative control molecules are provided for overexpression microRNA studies.
Successful microRNA functional studies begin with optimization of the assay in each cell line and/or type of interest. To optimize experimental conditions, we offer both housekeeping and endogenous positive controls for overexpression experiments using miRIDIAN Mimics.
Highlights
- Same structure and design as experimental miRIDIAN microRNA Mimics
- Target the 3' UTR (untranslated region) of the standard housekeeping genes, PPIB, or GAPDH (see Figure 1, Supporting Data)
- Allows a clean, straightforward, cleavage-based readout of mimic function
Applications
- Delivery and function can be evaluated to determine transfection conditions in each cell line/type
- Monitor the function of a mimic molecule at the mRNA level using standard transcript quantification methods, such as RT-qPCR
miRIDIAN microRNA Mimic Housekeeping Controls serve as optimization tools
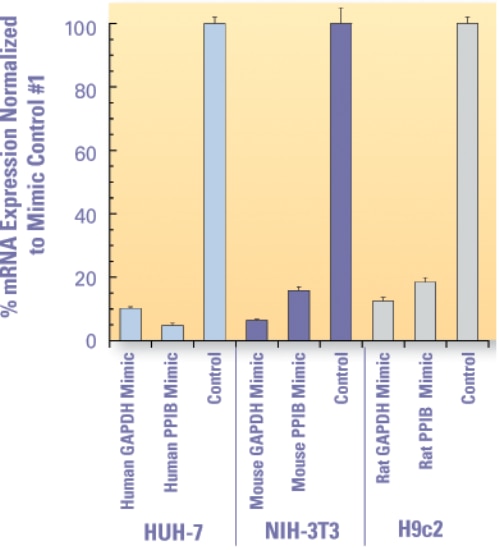
Figure 1. | miRIDIAN microRNA Mimics designed to target the 3' UTR of either PPIB or GAPDH were transfected at 50 nM using DharmaFECT 1 into the indicated cell lines and assessed for their ability to decrease target mRNA levels. PPIB or GAPDH down-regulation was determined using the Quantigene® branched DNA assay (Panomics) at 48 hours post-transfection.
Application notes
-
microRNA Mimic and Inhibitor Functional Analysis - Application Note
-
microRNA Targets in Stem Cell Differentiation - Application Note
-
Modulating endogenous miRNA targets with miRIDIAN microRNA mimics and inhibitors - Application Note
-
Screening microRNAs for Determinants of Osteogenesis - Application Note
Protocols
Related Products
The miRIDIAN microRNA Mimic Housekeeping Positive Controls allow for the direct monitoring of housekeeping genes in a microRNA mimic experiment. Positive Control #1 targets PPIB (aka Cyclophilin B).
The miRIDIAN microRNA Mimic Endogenous Positive Control helps you monitor specific mimic effects on target protein levels in a validated endogenous assay, which is based upon the targeted activity of miR-122 on Aldolase A mRNA levels in cell lines.
The miRIDIAN microRNA Mimic Transfection Control is a Cy3-labeled microRNA mimic based on the C. elegans miRNA cel-miR-67 (miRIDIAN Mimic Negative Control #1) for monitoring delivery into human, mouse, and rat cells.
Choose from two universal miRIDIAN Mimic Negative Controls for experiments with miRIDIAN microRNA Mimics. Negative control sequences based on C. elegans microRNAs have minimal sequence identity in human, mouse, and rat.
