- Gene modulation
- RNA interference
- siRNA solutions by Dharmacon™
- Accell siRNA Reagents
Accell siRNA Reagents
Guaranteed gene silencing in difficult-to-transfect cells
A novel siRNA for difficult-to-transfect cells, modified to require no transfection reagent or viral vector for delivery. Predesigned Accell siRNA is available as individual reagents and in SMARTpool format.

Accell siRNA Reagents
1Start Here
2Choose
Achieve RNAi in any cell type
Accell siRNA offers what no other RNAi product can claim: delivery into difficult-to-transfect cells without transfection reagents, virus, or instrumentation. Achieve targeted gene silencing in neuronal, immune, or primary cells that are beyond the reach of conventional RNAi products due to toxicity caused by transfection reagents or undesirable viral responses.
- Accell siRNA enters cells without the need for transfection reagents, virus (or viral vectors), or instruments
- Guaranteed silencing by SMARTpool and 3 of 4 individual siRNAs (see Guarantee tab)
- Novel siRNA modifications facilitate uptake, stability, specificity and knockdown efficiency
- Proven performance in neuronal, immunological, primary, and other difficult-to-transfect cell types
- Appropriate for use in vivo, due to stabilizing modifications
- Extended-duration knockdown with optimized continuous application
Experimental considerations
This breakthrough in siRNA delivery requires no transfection reagent, but has some unique application requirements.
- Accell siRNA works at a higher concentration than conventional siRNA; recommended 1µM working concentration
- Delivery may be inhibited by the presence of BSA in serum. Optimization studies with serum-free media formulations (Accell Delivery Media) or <2.5% serum in standard media is recommended
- Full-serum media can be added back after 48 hours of incubation, optimal mRNA silencing is typically achieved by 72 hoursor up to 96 hours for protein knockdown
Product Formats
- SMARTpool: A mixture of 4 siRNA provided as a single reagent,providing advantages in both potency and specificity
- Set of 4: A convenient option for purchasing aliquots of all 4 individual siRNAs targeting a single gene
- Individual siRNAs: Select 1, 2, 3, or 4 individual siRNAs per gene
Sequence information is provided with purchase of predesigned siRNA products.
Ordering Guidelines
| Approximate # reactions (wells) at 1µM siRNA concentration (assuming no loss from pipetting) | |||
|---|---|---|---|
| nmol | 96-well plate (100 µL total reaction volume) | 24-well plate (500 µL total reaction volume) | 12-well plate (1000 µL total reaction volume) |
| 1 | 10 | 2 | 1 |
| 2 | 20 | 4 | 2 |
| 5 | 50 | 10 | 5 |
| 10 | 100 | 20 | 10 |
Due to the unique nature of Accell siRNA delivery, it requires a higher working concentration than conventional siRNAs. The table provides the approximate number of reactions (wells) at recommended 1μM Accell siRNA working concentration in different plate formats.
Custom siRNA design
Design and order custom siRNA sequences using our siDESIGN center.
Our siRNA knockdown guarantee
Accell siRNA reagents (SMARTpool and three of four individual siRNAs) are guaranteed to silence target gene expression by at least 75% at the mRNA level.
If all of the steps to assess and optimize conditions are carried out as described in our "Guaranteed silencing with Accell siRNA in 3 easy steps" article, you are guaranteed at least 75% target gene silencing by the SMARTpool and/or three of the Set of 4 individual siRNAs. If, after contacting our scientific support team and carrying out recommended troubleshooting, your target gene is not silenced by at least 75%, a one-time replacement of the affected Accell SMARTpool or Set of 4 siRNA reagents will be provided.
Effectively optimize Accell siRNA delivery conditions and control for variables in ongoing experiments with fluorescent or unlabeled positive and negative controls. Accell Control Kits combine species-specific validated controls for assessment of Accell technology in your difficult-to-transfect cells. Kits of validated controls are available.
Experimental considerations
- Accell siRNA works at a higher concentration than conventional siRNA; recommended 1µM working concentration
- Delivery may be inhibited by the presence of BSA in serum. Optimization studies with serum-free media formulations (Accell Delivery Media) or <2.5% serum in standard media is recommended
- Full-serum media can be added back after 48 hours of incubation. Optimal mRNA silencing is typically achieved by 72 hours or up to 96 hours for protein knockdown
Choose from our selection of positive controls (targeting housekeeping genes), fluorescent or unlabeled negative controls, or an Accell Control Kit.
| Accell Positive Control Reagents | Species | Catalog Number |
|---|---|---|
| Accell Cyclophilin B Control siRNA | Human, Mouse, Rat | D-001920-0Y |
| Accell Cyclophilin B Control Pool | Human, Mouse, Rat | D-001920-Y0 |
| Accell GAPD Control siRNA | Human, Mouse, Rat | D-001930-0Y |
| Accell GAPD Control Pool | Human, Mouse, Rat | D-001930-Y0 |
| Accell Green Cyclophilin B Control siRNA | Human, Mouse, Rat | D-001970-0Y |
| Accell Red Cyclophilin B Control siRNA | Human, Mouse, Rat | D-001975-0Y |
| Accell eGFP Control siRNA | N/A | D-001940-01 |
| Accell eGFP Control Pool | N/A | D-001940-10 |
| Accell Negative Control Reagents | Species | Catalog Number |
| Accell Non-targeting siRNAs | Human, Mouse, Rat | D-001910-0X |
| Accell Non-targeting Pool | Human, Mouse, Rat | D-001910-10 |
| Accell Green Non-targeting siRNA | Human, Mouse, Rat | D-001950-01 |
| Accell Red Non-targeting siRNA | Human, Mouse, Rat | |
| Accell Control siRNA Kits | Species | Catalog Number |
| Accell Control siRNA Kits (Green) | Human, Mouse, Rat | K-005000-G1-0Y |
| Accell Control siRNA Kits (Red) | Human, Mouse, Rat | K-005000-R1-0Y |
Accell siRNA demonstrates effective uptake and silencing in organotypic brain slices
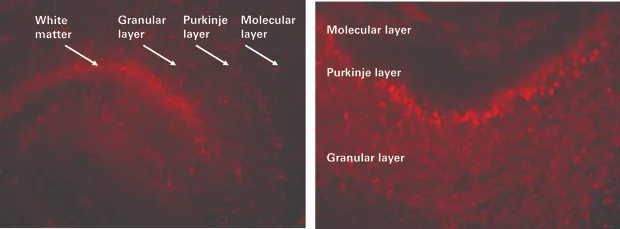
Accell siRNA shows increased uptake with extended incubation. 250 μm cerebellar sections were prepared, cultured, and incubated for 3 hours (A) and 72 hours (B) with Accell Red Non-targeting control (NTC) siRNA before inspection by microscopy.
Accell siRNA effectively silences target genes in cultured P8 mouse brain slices
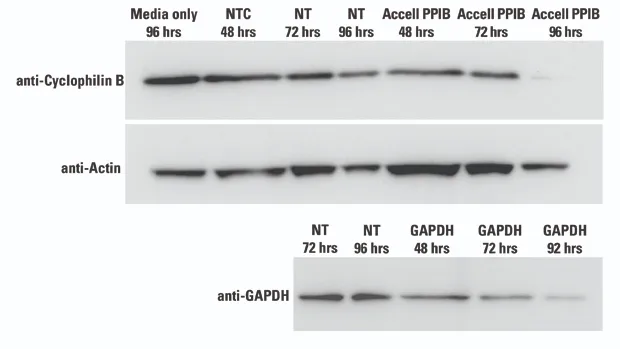
Cultured brain slices were assessed following incubation with Accell siRNA targeting PPIB and GAPDH or Non-targeting controls (NTC) at three time points (48, 72, 96 hours) and the level of remaining protein was evaluated using Western blot analysis (Actin as loading control).
Accell siRNA delivery and gene silencing in cardiomyocytes
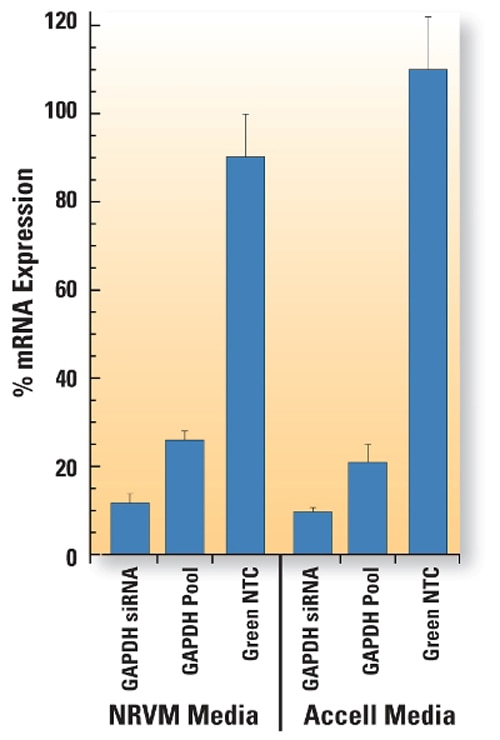
Neonatal rat ventricular myocytes were incubated with 1μM Accell Green (A; Cat# D-001950-01) or Red (B; Cat# D-001960-01) Non-targeting siRNA for 72 hours in Accell delivery media (Cat# B-005000). Nuclei were stained with DAPI (blue). Labeled control uptake showed diffuse cytoplasmic localization in nearly all cells. The bar graph indicates the level of gene silencing achieved with Accell GAPD Control siRNA (Cat# D-001930-03) and Accell GAPD Control Pool (Cat# D-001930-30) control reagents when used with neonatal rat ventricular myocyte (NRVM) media or Accell delivery media. Myocytes were prepared as described in Maass AH & Buvoli M. Cardiomyocyte preparation, culture, and gene transfer. Methods in Molecular Biology 2007;366: 321-30. mRNA expression was determined by QuantiGene branched DNA assay (Panomics).
The Accell siRNA application protocol simplifies targeted gene knockdown
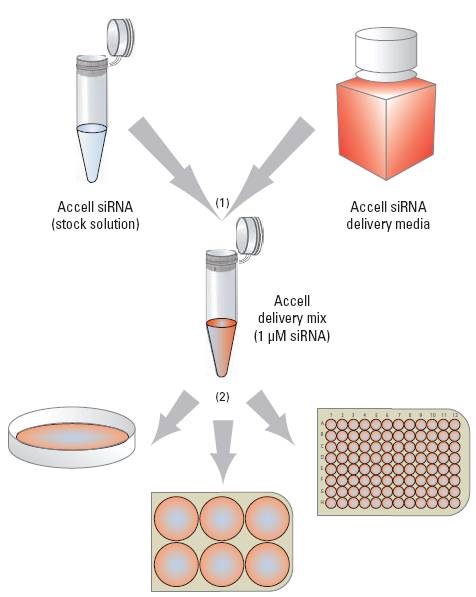
(1) Combine Accell siRNA with Accell delivery media (or other low- or no-serum media). (2) Add Accell delivery mix directly to cells, and incubate for 72 hours.
Application notes
Posters
Product data
Product inserts
Reading lists
Safety data sheets
Selection guides
Related Products
Validated positive control pool of four siRNAs targeting the Cyclophilin B (PPIB) housekeeping gene in human, mouse, or rat. Accell siRNA requires no transfection reagent or viral vector for delivery into difficult-to-transfect cells.
Catalog ID:D-001920-10-05
Unit Size:5 nmol
$269.00
Validated positive control pool of four siRNAs targeting the GAPD housekeeping gene in human, mouse, or rat. Accell siRNA requires no transfection reagent or viral vector for delivery into difficult-to-transfect cells.
Catalog ID:D-001930-10-05
Unit Size:5 nmol
$269.00
A negative control pool of four siRNAs to determine baseline cellular response to Accell siRNA application. Component siRNAs are designed to target no human, mouse, or rat genes.
Catalog ID:D-001910-10-05
Unit Size:5 nmol
$269.00
An enriched, serum-free media for use during Accell siRNA application. While not required for use with Accell siRNA, this media provides appropriate serum-free conditions for Accell uptake while maintaining cell health.
Catalog ID:B-005000-500
Unit Size:500 mL
