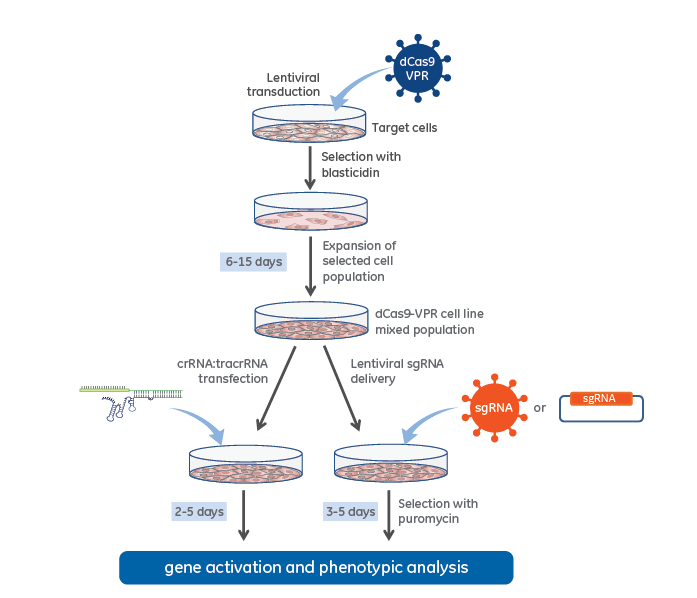
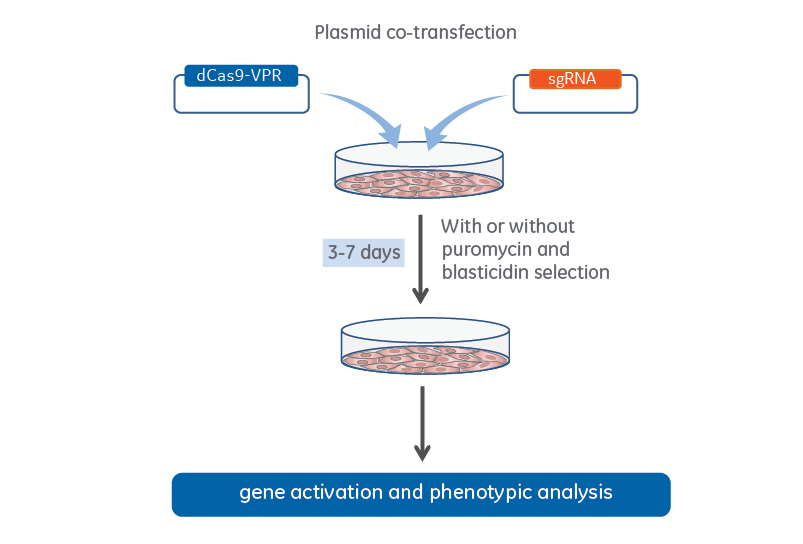
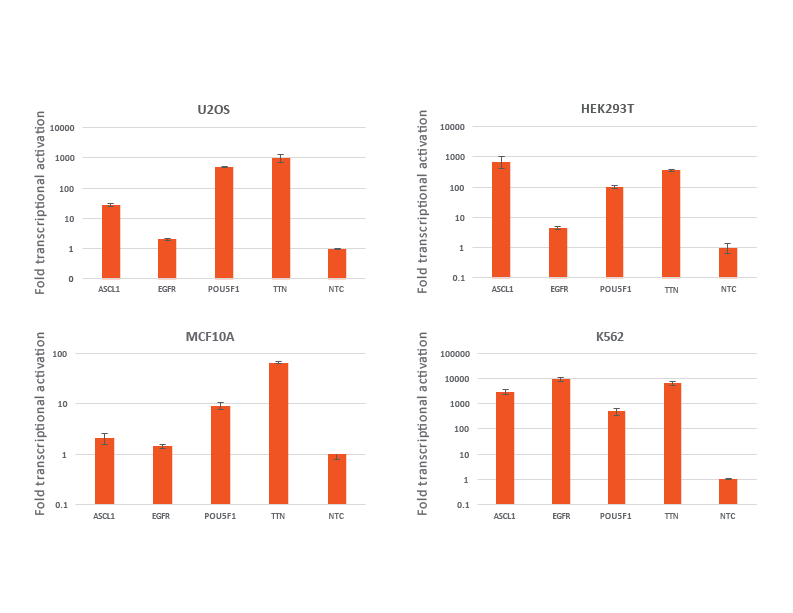
CRISPRmod CRISPRa lentiviral sgRNA non-targeting controls
Lentiviral sgRNA constructs bioinformatically designed and validated to not target any gene in human or mouse genomes
Negative control CRISPRa sgRNAs to establish experimental baselines and to distinguish sequence-specific biological effects from non-specific effects. Available as high-titer purified lentiviral particle and glycerol stock formats