Liquid biopsy is fast becoming a realistic clinical diagnostic tool, capable of profiling the molecular makeup of a tumor that aids in choosing the right drug and in monitoring treatment response. Studies for its use in early cancer detection are already underway with promising results1-3.
For liquid biopsy to become routine within the clinic, reliable, and affordable, diagnostic technologies, platforms and assay need to be readily available. This was a hot topic at AACR 2018, with the many cutting-edge companies showing promise in this area.
Horizon have developed and refined cell-free DNA (cfDNA) Reference Standards, which are an ideal companion for existing liquid biopsy assays. However, they are also a valuable tool for developing new technologies. As our material is cell line-derived, and highly characterized, it is ideal for validation studies to evidence reliability and reproducibility of novel technologies.
This case study, published in PLoS ONE4, explains how Horizon worked closely with Inivata, ctDNA and liquid biopsy specialists, to develop a new liquid biopsy platform.
Supporting the development of a new liquid biopsy platform
Inivata's enhanced TAm-Seq technology is an amplicon-based Next Generation Sequencing (NGS) method that can be used to identify clinically-relevant somatic alterations at low frequency in circulating tumor DNA (ctDNA). Using this sequencing technology, Inivata have developed a liquid biopsy platform - InVision - that can identify clinically-relevant somatic mutations at low frequency.
As with any new clinical diagnostic (CDx) technology, InVision required thorough validation throughout the development phases.
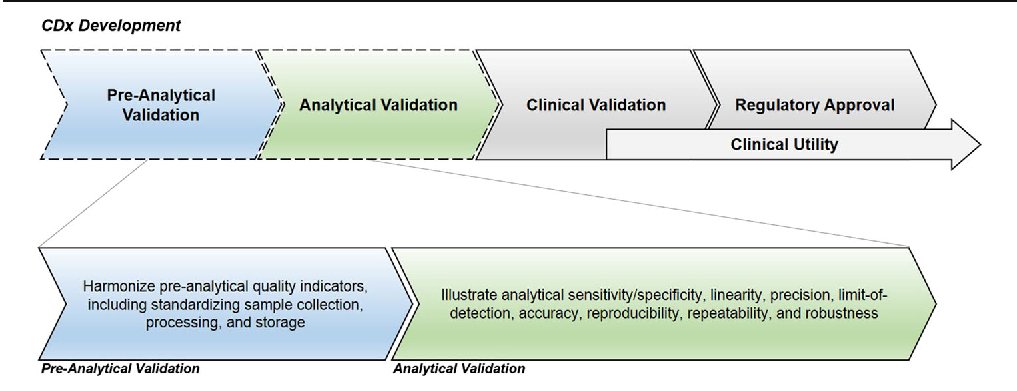
In the analytical validation stage of InVision's development, 3 separate studies were undertaken to evidence sensitivity and reproducibility using Horizon's Reference Standards:
Analytical validation of eTAm-Seq technology
To assess the sensitivity of this technology, analytical validation studies were performed in two labs. NGS libraries were prepared using eTAM-Seq, analyzing sheared reference standard DNA from Horizon and cfDNA extracted from control plasma.
Horizon provided two customized reference standards for Inivata for this study: Tru-Q6 and Tru-Q7, alongside the Tru-Q0 wild-type DNA, which were used as a dilutant. There are 21 mutations present in Tru-Q6 and 38 mutations in Tru-Q7 that were targeted by the InVision liquid biopsy tumor profiling panel.
Horizon's controls were also used to further assess the limit of detection, by creating a dilution series with sheared Tru-Q7 to an approximate AF range of 0.06% to 1.25% AF. Comparable results were seen across both laboratories and for all samples.
Comparison of performance of eTAm-Seq and digital PCR (dPCR) by analysis and novel full-process control material
Horizon cfDNA reference material was used to assess the quantitative performance of eTAm-Seq, by comparing the data with allele fractions generated by dPCR analysis of Horizon's Tru-Q6 and Tru-Q7. Results show good concordance between AFs determined by both technologies across 21 mutations present in both the InVision panel and Tru-Q6 and the 38 mutations common in Tru-Q7. As stated in the full article, this demonstrates the quantitative accuracy of eTAm-Seq technology for reliable detection of mutations at low allele frequency.
Analysis of novel full-process control material using eTAm-Seq technology and dPCR
Horizon's Multiplex I cfDNA reference standard was spiked-in to 16 presumed healthy donor samples. This standard contains 8 target mutations at 5%, 1% or 0.1% AF. By spiking into plasma containing background DNA, the mix would be expected to contain lower AFs than the original standards. Quantification of the 5% and 1% AF samples showed good concordance using both technologies.
The Results
This case study highlights how reference standards play a key role throughout the development of new sequencing technologies, and assays, and the versatility of our reference standards is showcased by their use to analyze sensitivity, quantitative accuracy and tested as a full-process control.
The results of these particular studies, alongside other tests undertaken during development, demonstrate that this new liquid biopsy suitable for the detection of low frequency clinically-relevant cancer mutations in cell-free DNA.
If you would like to see the full methods for each of these workflows, you can access the full article in our online resources, where you can also find other published work from Horizon and our partners.
References
- Santarpia M et al. Liquid biopsy for lunch cancer early detection. J Thorac Dis. 2018 April;10(Suppl 7):S882-S897
- Babayan A, Pantel K. Advances in liquid biopsy approaches for early detection and monitoring of cancer. Genome Med. 2018 Mar 20; 10(1):21
- Okada T et al. Utility of liquid biopsy using pancreatic juice for early detection of pancreatic cancer. Endosoc Int Open. 2018 Dec;6(12):E1454-E1461
- Gale D, Lawson ARJ, Howarth K, Madi M, Durham B, Smalley S, et al. Development of a highly sensitive liquid biopsy platform to detect clinically-relevant cancer mutations at low allelic fractions in cell-free DNA. PLoS ONE 13(3):e0194630
