The clinical success rate of new oncology drugs is only 3.4% compared to 20.9% in other disease types (Wong et al,2018). One contributing factor to this issue is the testing systems used, with two-dimensional (2D) monolayer assay formats as the traditional mainstay of high throughput screening. Although 2D monolayer assays have identified many successful drugs, it is increasingly recognized that they do not accurately model key aspects of the three-dimensional (3D) tumor environment. Therefore, the adoption of high throughput screening approaches using 3D assays to complement 2D approaches could substantially improve prediction of clinical outcomes and reduce the high failure rate of cancer drugs in clinical trials.
What’s involved with a spheroid-based high throughput 3D assay?
Many 3D assays have been described, but by their very nature most are typically complex often involving multiple cell types, extracellular scaffolds and complex image-based readouts: they are challenging to work with at scale. By contrast, 3D tumor spheroids produced by plating single cell suspensions of tumor cell lines in low anchorage U-bottom microplates in liquid growth media are amenable to straight forward add-and-read plate reader-based viability assays such as Promega’s Celltiter-Glo® 3D. This relative simplicity makes spheroid assays the most amenable to standardization for routine high throughput screening.
What can spheroids offer?
Increasingly, data indicate that tumor spheroid models are able to accurately mimic key aspects of the tumor microenvironment, including 3D cell-to-cell interactions and nutrient and oxygen gradients. They also uncover unique insights into drug responses that wouldn’t be discovered using traditional 2D assays. For example, Jeff Settleman and colleagues at Genetech elegantly applied both 3D spheroid-and standard 2D assays to identify a potential drug combination strategy to treat pancreatic cancer - a disease with a clear unmet clinical need. 'Using a two-dimensional monolayer culture system as well as three-dimensional spheroid culture system, we conducted a screen of a large panel of anticancer agents. These results reveal a combination drug treatment that might be effective in pancreatic cancer' (Sahu et al, 2017).
These data and our own in house data support the hypothesis that 3D drug panel screening could help to help predict which patient populations might be most responsive to a drug or biologic. Thus, Horizon has established a panel of approximately 200 cell lines to screen single agents and drug combinations using high throughput assays (Scales et al,2018a).
Making it more real - Organoid-based high throughput 3D assays
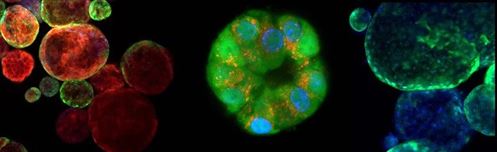
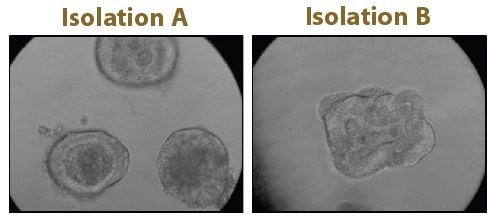
Although spheroids derived from cancer cell lines are an excellent tool for 3D screens, they lack the cellular heterogeneity of the originating tumor. This limitation can be addressed through patient-derived organoids, identified as Method of the Year by Nature Methods in 2017. These organoids are heterogenous and better represent the microanatomy of the original tumor, see Figure 1. Challenges that have limited the use of organoids for HTS, such as limited quantities of material and significant batch-to-batch variation, are being met by companies, such as Cellesce, through innovative bioprocessing technologies. In order to offer an organoid-based HTS service to clients, Horizon is developing robust and reproducible organoid-based assays, using colorectal cancer organoids generated by Cellesce (Scales et al,2018b).All these developments are making the path from research to therapy more straightforward.
References
- Sahu, N et al (2017) Cotargeting of MEK and PDGFR/STAT3 pathways to treat pancreatic ductal adenocarcinoma. Molecular Cancer Therapeutics, Vol 16(9), pp1729-1738.
- Scales, T et al (2018) High-Throughput cell panel and organoid screening in 3D.Poster presentation, ELRIG London, October 2018.
- Scales, T el al (2018) High-Throughput screening in colorectal organoids - proof of concept study. Horizon Application note 2018.
- Wong, C et al (2019) Estimation of clinical trial success rates and related parameters. Biostatistics Vol 20 (2), pp273-286.
