Getting a long list of potential genes that influence the response to your new therapeutic is one sign of a successful CRISPR screen, but the hard work has only just begun. How do you prioritize this list for targets that could have an impact in the clinic? Bioinformatics and data mining can help, but other screens can also add value, including large cell panel screens with the same therapeutic of interest.
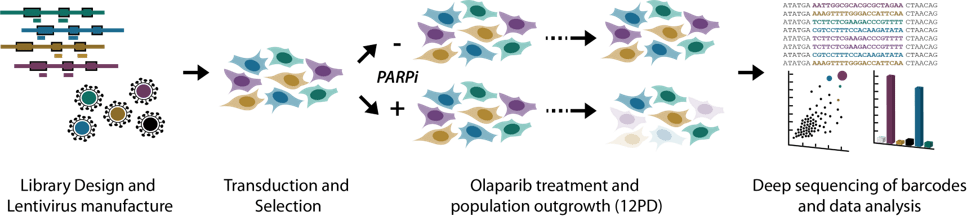
Pooled Screening workflow
A flexible CRISPRko library to fit any workflow
Horizon scientists have recently built a new, whole genome CRISPR knockout library that enables the use of 4, 6 or 8 guides, depending on the biological question being addressed and the modality being used for the screen. For example, eight guides could be used to find genes that when knocked out result in sensitivity to a new drug in cancer cell lines in culture (a drop out screen), whereas a four guide library might be best placed for in vivo CDX based CRISPR screens where there is a need to keep the library small.
A proof-of-principle screen
To test the utility of this new CRISPRko library, a whole genome drug–gene interaction screen was carried out in the HT 29 cancer cell line using the PARP inhibitor olaparib. The screen successfully identified genes that when knocked out either increased or decreased sensitivity to the drug compared with DMSO treated control cells.
So far, so good, but how best to triage this list?
Large cell panel screens, using many cancer cell lines representing different cancer types, can also be informative in identifying sensitivity and resistance mechanisms. Horizon scientists tested olaparib and an additional PARP inhibitor (talazoparib) across a panel of more than 300 cancer cell lines. Using these data and drug response and CRISPR screen data from the DepMap portal, a set of genes influencing sensitivity and resistance to these drugs was also compiled.
Substantial "hit" overlap with complementary methods
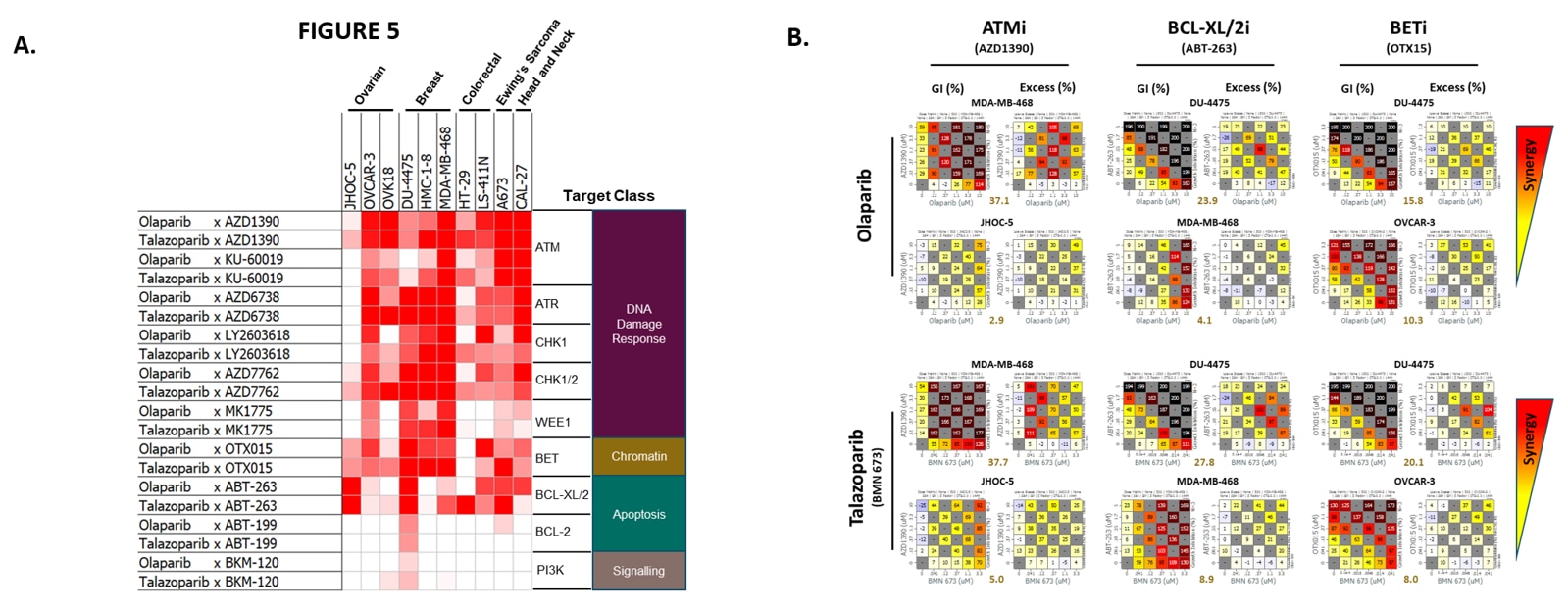
The genes influencing sensitivity and resistance to olaparib and talazoparib in the cell panel screen had substantial overlap with the genes identified in the CRISPR screen. Genes involved in DNA damage response, cell survival and cell division pathways featured heavily. A combination drug screen using clinical grade compounds was used to initially validate some of the genes whose loss seems to increase sensitivity to olaparib. Drug-mediated inhibition of proteins involved in DNA damage, such as ATM, ATR, CHK1 and CHK2; proteins involved in cell survival, such as BCL-XL; and proteins involved in cell division, such as WEE1, all increased sensitivity to olaparib.
Our integrated pooled CRISPRko and high throughput cell panel screening platform offers more clinically relevant information
Assessing overall concordance between our CRISPRko screen in HT29 cells and similar CRISPRko screens published in other cell types using olaparib shows that agreement between CRISPR screens carried out in different labs is substantially greater than that observed in RNA interference screens. However, differences between cell lines are still evident, indicating that robust conclusions from drug-gene interaction screens are best made from screens conducted across diverse genetic and lineage backgrounds, to better describe the drug–gene effects. This can be achieved with our fully integrated pooled CRISPRko and high throughput cell panel screening platform. The orthogonal approach of cell panel screening delivers rapid initial validation of selective hits in high throughput and enables targets to be considered in terms of their tractability in the clinic. These data also illustrate how CRISPR screens and high throughput cell panel screens can be used to rapidly validate new, clinically relevant genetic interactions.
AB Blanck 2020
 Written by Dr. Nicola McCarthy
Written by Dr. Nicola McCarthy
Dr, McCarthy has 10 years of postdoctoral research experience studying apoptosis (programmed cell death) in the context of cancer research and cardiovascular research and studied human anatomy for her B.Sc. and programmed cell death for her Ph.D. at the University of Birmingham, UK.
Read the full publication in The CRISPR Journal
A flexible, pooled CRISPR library for drug development screens. Blanck, M., et al. CRISPR J. 3: 211-222 2020. doi: 10.1089/crispr.2019.0066
