- Gene modulation
- RNA interference
- siRNA solutions by Dharmacon™
- siGENOME siRNA Reagents
siGENOME siRNA Reagents
Trusted for high-confidence silencing since 2002

siGENOME siRNA Reagents
1Start Here
2Choose
Straightforward and reliable gene knockdown
siGENOME siRNA reagents are widely recognized and trusted for highly efficient target gene silencing. Our scientists were the first to establish siRNA design rules for high potency silencing and continue to drive innovation in this area. By providing genome-wide siRNAs designed with the SMARTselection algorithm, researchers can focus on successful RNAi experiments instead of optimizing siRNA designs for their genes of interest.
siGENOME siRNA is available as pre-designed, genome-wide SMARTpool and individual siRNA reagents for human, mouse and rat. Simply search for your gene of interest and choose from the available product formats and quantities.
- Guaranteed knockdown by SMARTpool and 3 of 4 individual siRNAs (see Guarantee tab)
- Antisense strand loading into RISC ensured by thermodynamic analysis and selective application of a sense strand-blocking modification
- Sequence information provided with siRNA purchase
Design and modification strategies for optimal silencing
In addition to identifying potent siRNA designs with the SMARTselection algorithm, the creation of siGENOME siRNAs also involves:
- Thermodynamic analysis to determine optimal antisense strand-loading characteristics for effective silencing
- BLAST analysis of both strands to alleviate cross-targeting
To retain more high-scoring siRNA designs, the above analysis may trigger the use of a sense strand-blocking modification to ensure only the appropriate strand enters RISC for effective target knockdown (see Figure 2, Supporting Data).
Product Formats
- SMARTpool: A mixture of 4 siRNA provided as a single reagent; providing advantages in both potency and specificity.
- Set of 4: A convenient option for purchasing aliquots of all 4 individual siRNAs targeting a single gene.
- Individual siRNAs: Select 1, 2, 3 or 4 individual siRNAs per gene.
Order Quantity Guidelines
siGENOME reagents are routinely used at 5 to 25 nM concentration. The calculations below, based on 25 nM, are for estimation purposes only and assume no loss from pipetting.
| Approximate # reactions (wells) at 25 nM siRNA concentration | |||
|---|---|---|---|
| nmol | 96-well plate(100 µL total reaction volume) | 24-well plate(500 µL total reaction volume) | 12-well plate(1000 µL total reaction volume) |
| 1 | 400 | 80 | 40 |
| 2 | 800 | 160 | 80 |
| 5 | 2000 | 400 | 200 |
| 10 | 4000 | 800 | 400 |
| 20 | 8000 | 1600 | 800 |
Custom siRNA design
Design and order custom siRNA sequences using our siDESIGN center.
Our siRNA knockdown guarantee
siGENOME siRNA reagents (SMARTpool and three of four individual siRNAs) are guaranteed to silence target gene expression by at least 75% at the mRNA level when demonstrated to have been used under optimal delivery conditions (confirmed using validated positive control and measured at the mRNA level 24 to 48 hours after transfection using 100 nM siRNA).
Note: Most siGENOME siRNA products are highly functional at 5 to 25 nM working concentration.
A robust selection of validated positive, negative, and novel RISC-Free® controls to effectively assess all aspects of RNAi experiments. Controls are critical for the successful application of RNAi. Including the appropriate controls in each experiment adds certainty to the cause-and-effect relationship between gene silencing and siRNA, and aids in the distinction between target-specific and potential off-target effects.
Select a species-specific positive control to silence an expressed housekeeping gene in your experimental cell type. Our panel of non-targeting controls permits assessment of potential non-specific effects, to find the optimal negative control for your cell type, and your assay.
| siGENOME Positive Control Reagents | Species | Catalog Number |
|---|---|---|
| siGENOME Cyclophilin B Control siRNA | Human, Mouse, Rat | D-001136-01 |
| siGENOME GAPD Control siRNA | Human | D-001140-01 |
| siGENOME Lamin A/C Control siRNA | Human, Mouse, Rat | D-001050-01 |
| siGENOME Negative Control Reagents | Species | Catalog Number |
| siGENOME Non-targeting siRNAs | Human, Mouse, Rat | D-001210-0X |
| siGENOME Non-targeting siRNA Pools | Human, Mouse, Rat | D-001206-13 |
| siGENOME RISC-Free siRNA | Human, Mouse, Rat | D-001220-01 |
| siGENOME Control siRNA Kits | Species | Catalog Number |
| siGENOME Controls Basic Kit | Human, Mouse, Rat | K-002800-C4-01 |
| siGENOME Controls Complete Kit | Human, Mouse, Rat | K-002800-C2-0X |
siGENOME SMARTpool reagents demonstrate potency at 5 nM working concentration
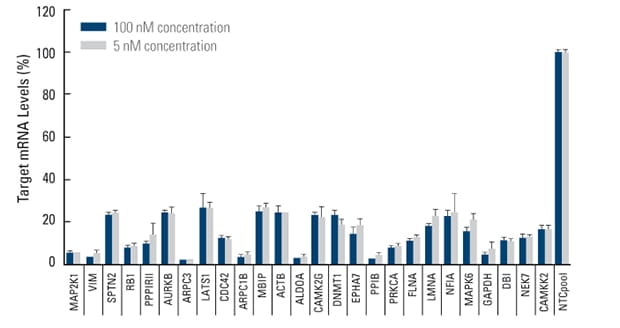
A comparison of target gene silencing at 5 and 100 nM concentration illustrates the high potency of siGENOME SMARTpool reagents. 5 or 100 nM siGENOME SMARTpool siRNA reagents targeting the indicated genes was transfected into HeLa cells (10K cells/well) using 0.1 μL/well DharmaFECT 1. The negative control (NTC pool) was siGENOME Non-Targeting Pool #2.
Unnecessary sense-strand inactivation can increase off-target activity
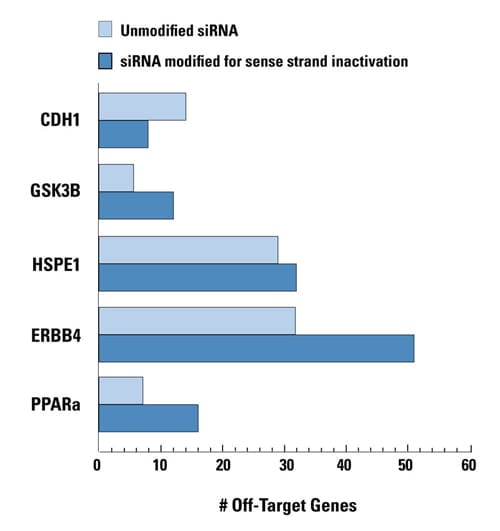
Unmodified and sense strand-inactivated siRNAs were used to target five genes. In four cases off-targets were increased due to enhanced RISC loading of the antisense strand when the sense strand was modified. The unmodified siRNAs have natural guide-strand loading characteristics. All siRNAs had comparable silencing potency. Data shown represents genes down-regulated by twofold or more. HEK293 cells were transfected with 100 nM siRNA using 0.2 μL of DharmaFECT 1. Data was analyzed at 24 hours by genomewide microarray analysis (Agilent).
Why not modify ALL siGENOME siRNAs to ensure proper strand loading? It has been demonstrated by Dharmacon scientists1 and others2 that forcing antisense (guide) strand entry into RISC may actually INCREASE off-targets due to increased loading of the guide strand and resulting off-target activity by its seed region. siGENOME siRNAs are designed with thermodynamic properties to naturally facilitate guide strand entry to RISC, which has been demonstrated to correlate with functionality1. However, in cases where a high-scoring siGENOME siRNA does not possess ideal strand-loading characteristics, a sense (passenger) strand-inhibiting chemical modification (ON-TARGET) is utilized to promote guide strand entry. Approximately 20% of siGENOME siRNAs carry the ON-TARGET modification.
Microarray analysis demonstrates SMARTpool potency and reduced off-targets
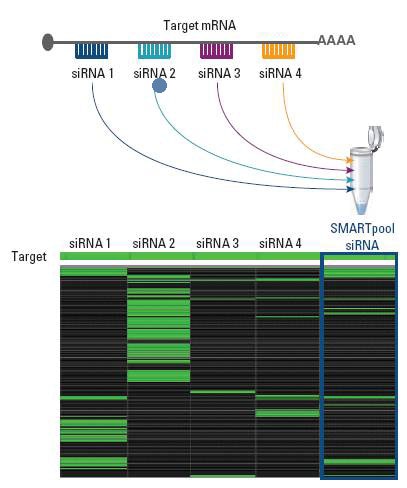
Target gene silencing is maintained while off-target signatures are reduced with a SMARTpool siRNA reagent when compared to its four constituent siRNAs.
Total RNA for microarray extracted from HeLa cells 24 hours post-transfection. Green indicates mRNA reduced by two-fold or more (Agilent 22K Platform).- A. Reynolds et al., Rational siRNA design for RNA interference. Nature Biotechnology.22(3), 326-30 (March 2004).
- P. Y. Chenet al., Strand specific 5' O-methylation of siRNA duplexes controls guide strand selection and targeting specificity. RNA.14(2), 263-274 (February 2008).
Application notes
Product inserts
Protocols
Safety data sheets
Selection guides
Related Products
Concentrated buffer solution recommended for resuspension and long-term storage of any short, double-strand, or single-strand synthetic RNA molecule. Dilute with RNase-free water prior to use.
Molecular grade water for dilution of 5x siRNA Buffer or resuspension of RNA. RNase-free to prevent degradation of RNA reagents and oligonucleotides.
The most broadly applicable DharmaFECT formulation for optimal siRNA or microRNA transfection into a wide range of cell types for successful RNAi experiments
Catalog ID:T-2001-01
Unit Size:0.2 mL
$116.00
A validated positive silencing control targeting the Cyclophilin B (PPIB) gene in human, mouse, or rat cell lines. Useful for determination of optimal RNAi experimental conditions
Catalog ID:D-001136-01-05
Unit Size:5 nmol

