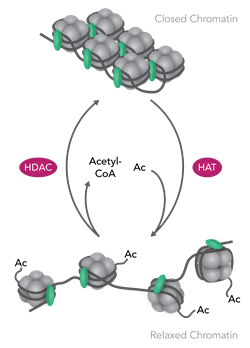
One key modification is histone acetylation of the N-terminal tail and this is regulated by histone acetyl transferases (HATs) and histone deacetylases (HDACs). Acetylation is a type of epigenetic marker of chromatin and results in the condensed chromatin “relaxing” and in this state it is known as euchromatin, this allows greater access to the DNA for proteins such as transcription factors and results in greater levels of gene transcription.
Histone Acetylation Imbalance
HDACs are enzymes which reverse the acetylation of histones, resulting in tightening of the chromatin (heterochromatin) and decreases in gene transcription. Imbalance of histone acetylation is a common aspect of many disorders and HDAC inhibitors are of interest for treatment of cancers, neurodegenerative diseases and metabolic disorders.
Cell Lines for Studying Histone Acetylation
|
clone 1013-10 |
Abcam ab187947 |
|
|
clone 1012-4 |
Abcam ab187941 |
|
|
clone 579-3 |
Abcam Ab7030 |
|
|
clone 1015-10 |
Santa Cruz sc11405 |
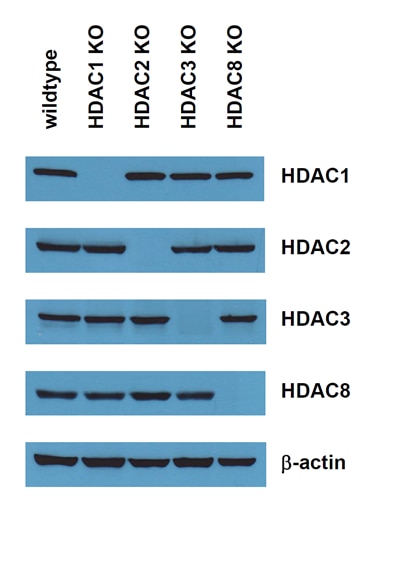
Examples of popular knockout cell lines for Histone Acetylation
|
|
|||
|
|
|||
|
|
Order products
Human knockout HAP-1 cells
The single largest bank of isogenic cell lines with over 7,500 cell lines to choose from and trusted by academia, biotech, and pharma research labs.
Cancer-related cell lines
Choose from over 300 knock-in and knockout cell line models in many standard cancer cell lines such as DLD1, MCF10A, and HCT116.
Cas9 Stable Cell Lines
Simplify gene editing experiments with stably expressing Cas9 cell lines
CRISPRmod CRISPRa dCas9-VPR Stable Cell Lines
Streamline CRISPR activation experiments with stably expressing dCas9-VPR cell lines
