- Gene editing
- Gene editing reagents
- Edit-R predesigned synthetic sgRNA
Edit-R predesigned synthetic sgRNA
Synthetic single guide RNAs for efficient gene knockout & unparalleled specificity
- Guaranteed to edit the target gene of interest
- Algorithm-optimized to maximize functional protein knockout and minimize off-target editing
- Transfection-ready synthetic RNAs eliminate cloning and in vitro transcription steps
- Chemically modified for improved nuclease resistance
- Edit-R human sgRNA designs have been updated to the latest RefSeq as of August 22, 2025.
- Pair your predesigned synthetic sgRNA with a Cas9 nuclease including our New! Edit-R Cas9 Protein Hybrid NLS

Edit-R predesigned synthetic sgRNA
1Start Here
2Choose
Functional and specific targeting for high-confidence gene knockout results
Edit-R predesigned synthetic sgRNAs are chemically synthesized guide RNA for S. pyogenes Cas9 nuclease targeting and creation of Cas9-mediated DNA double-strand breaks in a genomic region of interest.
All Edit-R guides are algorithmically designed to efficiently generate functional protein knockouts, not just create an insertion or deletion. Additionally, chemical modifications are applied to all Edit-R synthetic guide RNA products to reduce degradation by nucleases and improve overall editing performance.
This unmatched accuracy and efficiency makes Edit-R predesigned synthetic sgRNA an ideal tool for pathway analysis or following up hits from a synthetic sgRNA library screen.
New! Edit-R human sgRNA designs have been updated to the latest RefSeq in 2025 providing the most specific and genomically relevant guides for producing efficient protein knockout. This allows the Edit-R algorithm to target the latest genome annotations more accurately and efficiently providing you with the best solution for your research needs. Please reach out to Scientific Support if you have any questions or read our blog on revvity.com.
All guide RNA designs are top algorithm picks for each gene; qualitative ranks for functionality and specificity allow you to fine tune human guide RNA choice to your specific application. The functionality score is a predicted indication of how likely this guide is to produce a functional knockout. The specificity score is based on the predicted risk of cutting activity at potential off-target sites. To learn more, visit our algorithm for Edit-R guide RNA page
Components for CRISPR gene editing experiment using synthetic sgRNA
- Edit-R synthetic sgRNA designed to target the gene of interest — choose from individual guides or sets of 3 guides for greater likelihood of functional knockdown
- A source of Cas9 nuclease — choose from protein, mRNA, or lentivirus formats, or a stable Cas9 expressing cell line
- Appropriate positive and non-targeting controls
- DharmaFECT transfection reagent or electroporation for guide delivery
Need to scale up your experiments? Choose from arrayed collections of predesigned synthetic sgRNA libraries for screening across the human genome and gene families, or design your own custom mini-library with our Cherry-Pick library tool.
The Edit-R guide RNA guarantee
We guarantee that EVERY predesigned Edit-R guide RNA will provide successful editing at the target site when delivered as described in the Edit-R Technical Manuals.
The Edit-R guide RNA guarantee is valid when used with any wild type S. pyogenes Cas9 nuclease, including mRNA, expression plasmid, protein, or stable Cas9 expression.
Analysis of editing of the treated cell population must be shown using a T7EI or Surveyor mismatch detection assay. If successful editing is not observed for a predesigned Edit-R guide RNA while an appropriate side-by-side Edit-R positive control is successful, a one-time replacement of a different predesigned Edit-R guide RNA of the same format and quantity will be provided at no cost.
A replacement will only be approved upon discussion with our Scientific Support team.
Successful editing at the DNA level does not always lead to functional gene knockout; it is recommended to test multiple guide RNAs to determine the most effective guide RNA for knockout of your target gene.
This guarantee does not extend to any accompanying experimental costs, does not apply to guide RNAs ordered via the CRISPR Design Tool, and will not be extended to the replacement guide RNA.
Gene knockout workflow using Cas9 nuclease mRNA and synthetic sgRNA or crRNA:tracrRNA
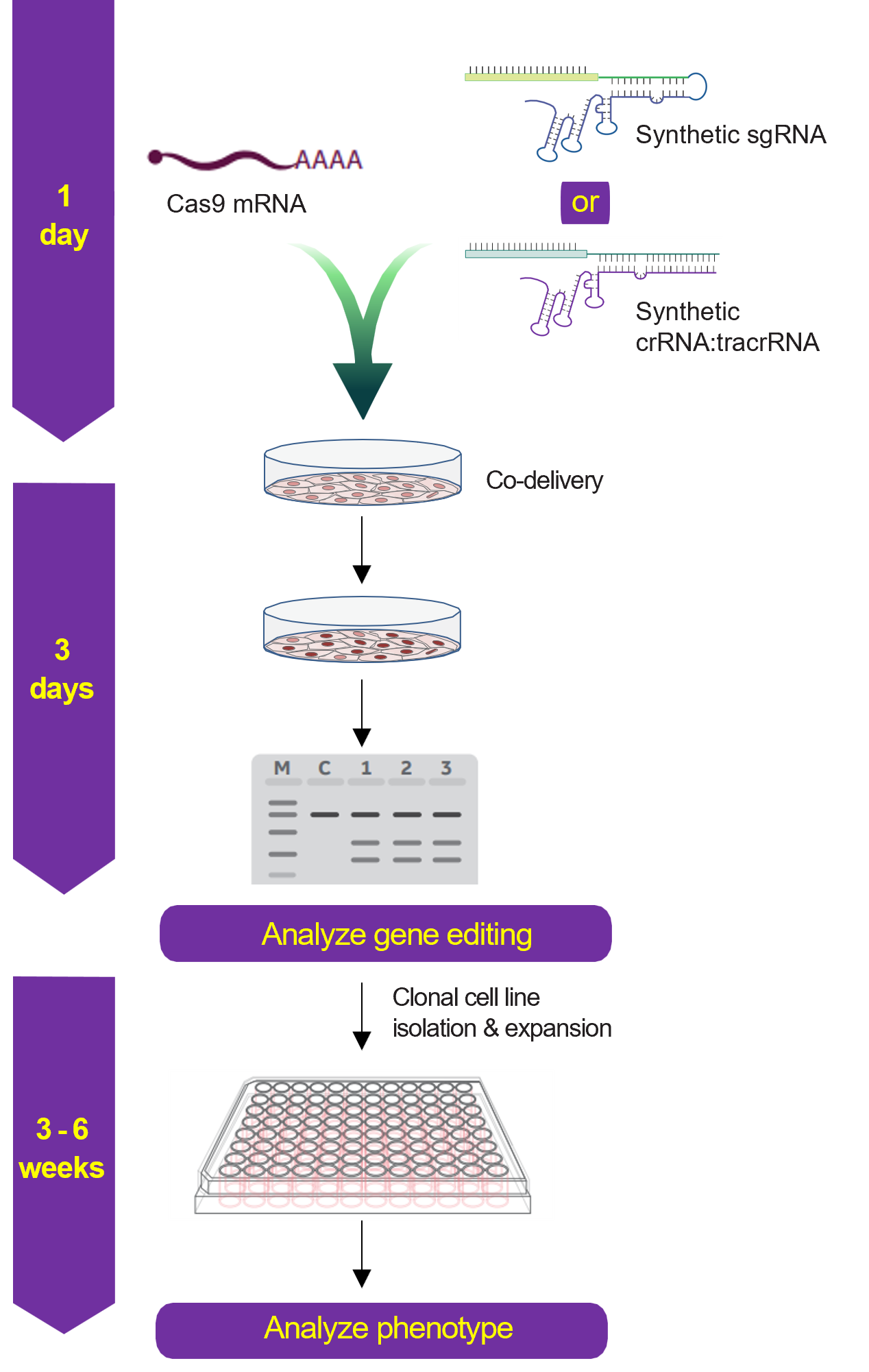
Gene knockout workflow using Cas9 nuclease protein and synthetic sgRNA or crRNA:tracrRNA

Gene knockout workflow using lentiviral Cas9 expression particles and synthetic sgRNA or crRNA:tracrRNA

Complete your knockout experiment by adding these supporting reagents to your order
Cas9 nuclease options
- Cas9 mRNA or Cas9 protein NLS can be co-transfected with Edit-R synthetic sgRNA for a fully DNA-free workflow.
- Stable Cas9 expressing cell lines are ready to be transfected with Edit-R synthetic sgRNA for rapid gene knockout.
Synthetic guide RNA controls
- Non-targeting controls are used to evaluate cellular responses to CRISPR-Cas9 components in the absence of gene target-specific sgRNA. Choose from 5 scrambled sequences guaranteed not to target any gene in the human or mouse genomes.
-
Cutting controls
Cutting control (safe harbor) synthetic sgRNAs recommended for determination of baseline cellular responses in CRISPR-Cas9 experiments.
- Positive controls targeting well-characterized genes are used to determine the effectiveness of your gene editing conditions for maximal efficiency. Choose from PPIB or DNMT3B.
Transfection reagents
- DharmaFECT transfection reagents are optimized for improved delivery and reduced toxicity. The optimal DharmaFECT reagent for your experiment will depend on your Cas9 nuclease source and your cell type.
| Cas9 nuclease source | Recommended transfection reagent |
|---|---|
| Stable Cas9 expressing cell line or Cas9 nuclease lentiviral particles | DharmaFECT 1,2,3, or 4 for transfection of synthetic guide RNA. Use this guide to find the recommended DharmaFECT reagent for your cell type. |
| Cas9 mRNA or protein NLS | DharmaFECT Duo for co-transfection of mRNA or protein and synthetic guide RNA |
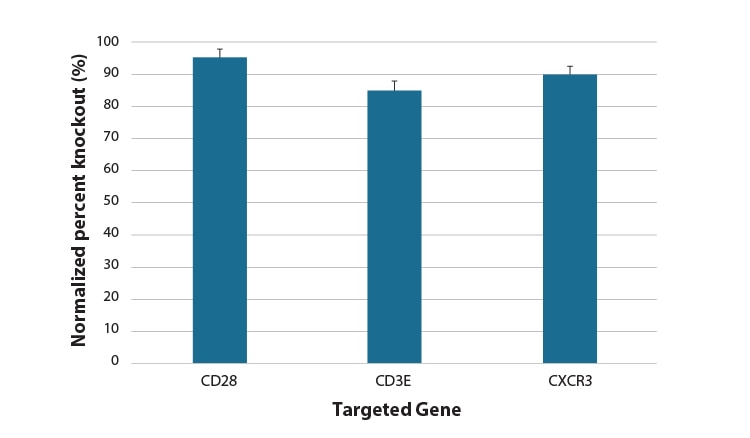
Flow cytometry analysis of protein knockout in CD4+ T cells using a single predesigned synthetic sgRNA per gene target
 |
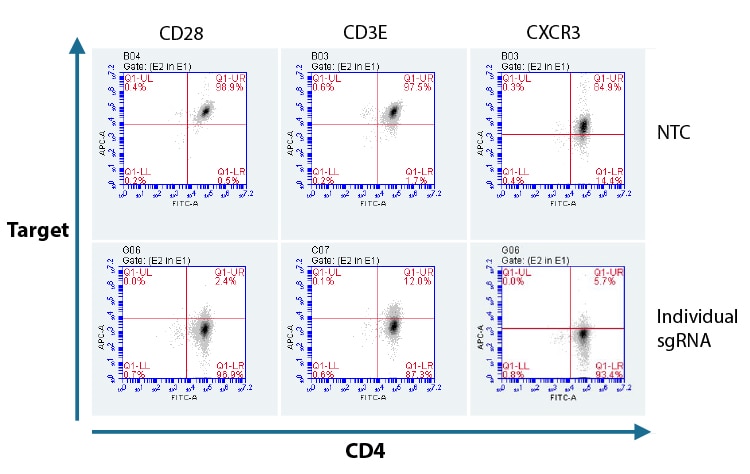 |
Primary human CD4+ T cells were nucleofected with Cas9 RNP using an individual predesigned synthetic sgRNA per gene target or a non-targeting control (NTC), via a Lonza 96-well Shuttle system. After 72 hours, functional knockout of CD28, CD3E, and CXCR3 were assessed as a percent of cells not expressing the target gene by FACS analysis. Cells were stained and normalized for CD4 expression using an Alexa Fluor 488 conjugated antibody and compared to each target CD28, CD3E, and CXCR3 using APC conjugated primary antibodies.
Functional knockout of FUS gene with a pool of synthetic sgRNA
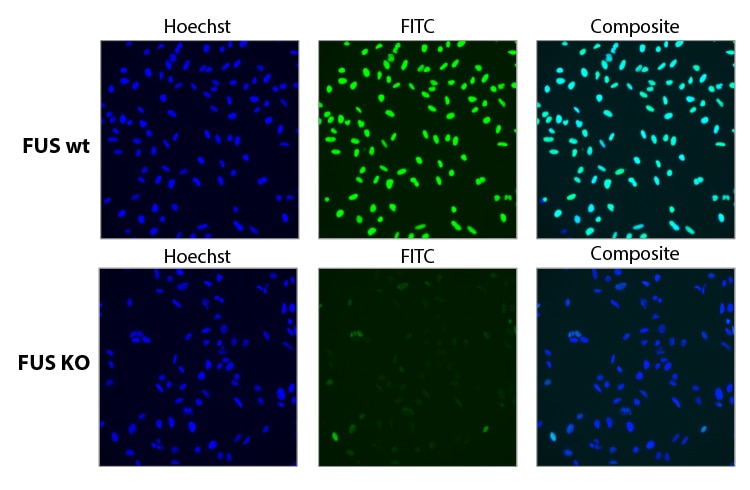
Functional knockout of FUS was assessed in U2OS cells constitutively expressing Cas9 under the CAG promoter. Cells were transfected with a 25 nM sgRNA pool containing three different Edit-R predesigned sgRNAs targeting FUS using 0.04 µL DharmaFECT 4. 72 hours post transfection the cells were split, and at 96 hours post transfection, cells were fixed and stained with a primary antibody targeting FUS, and an Alexa Fluor 488 conjugated fluorescent secondary antibody. Hoechst stain was used to identify nuclei.
Nuclease stabilization modifications improve gene editing efficiency
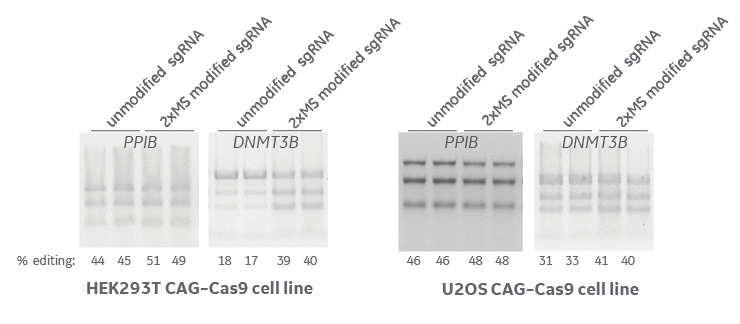
HEK293T and U2OS integrated Cas9 (under the CAG promoter) cell lines were transfected with unmodified and 5’ and 3’ 2xMS modified synthetic sgRNA targeting PPIB and DNMT3B genes. At 72 hours editing efficiency was assessed with a mismatch detection assay, T7 Endonuclease I (T7EI). In all cases the stabilizing modifications improve gene editing efficiency. MS = 2’-O-methyl nucleotides and phosphorothioate backbone linkage.
Edit-R synthetic guide RNAs cause virtually no innate immune response or toxicity compared to in vitro transcribed guide RNA

A HEK293T Cas9 nuclease expressing cell line was transfected with different synthetic guide RNA formats including unmodified crRNA:tracrRNA, crRNA:tracrRNA modified with 2xMS on 5’ crRNA and 3’ tracrRNA, crRNA:tracrRNA modified with 3xMS on both 5’ and 3’ crRNA and tracrRNA, unmodified synthetic sgRNA, modified synthetic sgRNA with 5’ and 3’ 2xMS or 3xMS, and in vitro transcribed (IVT) sgRNA targeting PPIB and DNMT3B genes. At 72 hours viability was assessed with the Resazurin reduction assay (red dots) and the levels of five immune response genes were quantified by RT-qPCR. MS = 2’-O-methyl nucleotides and phosphorothioate backbone linkages.
Structure of modifications for improved nuclease resistance on Edit-R synthetic sgRNA
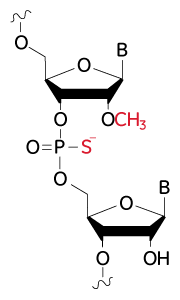
Modifications for improved nuclease resistance: 2’-O-methyl modified nucleotides and phosphorothioate backbone linkages.
Sequence structure of Edit-R synthetic single guide RNA
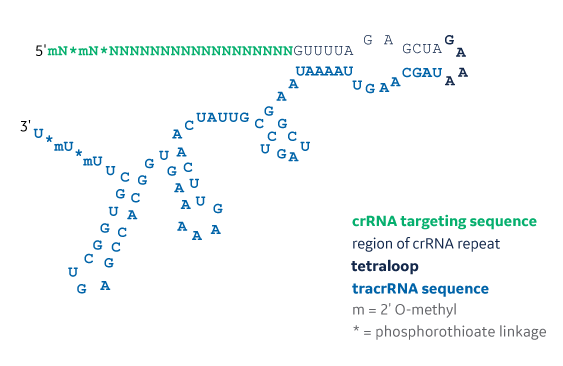
The structure of a Edit-R synthetic single guide RNA showing the 2xMS nuclease stability modifications on both 5’ and 3’ ends of the molecule. The structure contains a 20 nt targeting sequence (shown in green as poly Ns), 12 nt crRNA repeat sequence (shown in light gray), 4 nt tetraloop sequence (shown in bold black), and a 64 nt tracrRNA sequence (shown in blue). MS = 2’O-methyl nucleotides and phosphorothioate linkages (MS).
How much sgRNA do I need?
This table provides the approximate number of experiments that can be carried out for lipid transfection methods at the recommended sgRNA working concentration (25nM) in various plate/well formats. Calculations do not account for pipetting errors.
| sgRNA nmol | 96-well plate 100 µL reaction volume | 24-well plate 500 µL reaction volume | 12-well plate 1000 µL reaction volume | 6-well plate 2500 µL reaction volume |
|---|---|---|---|---|
| 2 | 800 | 160 | 80 | 32 |
| 5 | 2000 | 400 | 200 | 80 |
| 10 | 4000 | 800 | 400 | 160 |
- R. Barrangou, A. Birmingham, et. al. Advances in CRISPR-Cas9 genome engineering: lessons learned from RNA interference. Nucleic Acids Research, 43(7) 3407-3419 (2015)
- M.L. Kelley, Ž. Strezoska, et al. Versatility of chemically synthesized guide RNAs for CRISPR-Cas9 genome editing. J. Biotechnol. 233, 74–83 (2016). doi:10.1016/j.jbiotec.2016.06.011
- Basila, M., M. L. Kelley, et al.. Minimal 2’-O-methyl phosphorothioate linkage modification pattern of synthetic guide RNAs for increased stability and efficient CRISPR-Cas9 gene editing avoiding cellular toxicity. PLoS One. 12, e0188593 (2017). doi: 10.1371/journal.pone.0188593
- Anderson, E.M., A. Haupt, et al. Systematic analysis of CRISPR-Cas9 mismatch tolerance reveals low levels of off-target activity. J. Biotechnol. 211, 56-65 (2015)
- He, K., E. Chou, et al. Conjugation and evaluation of triazole-linked single guide RNA for CRISPR-Cas9 gene editing. ChemBioChem. DOI: 10.1002/cbic.201600320 (2016)
Application notes
Product data
Protocols
-
Cas9 Nuclease mRNA Electroporation - Protocol
-
Cas9 nuclease protein and Edit-R synthetic guide RNA transfection - Quick Protocol
-
Cas9 nuclease protein electroporation - Protocol
-
Synthetic guide RNA resuspension - Protocol
-
T7E1, TIDE, and NGS analysis protocol for Dharmacon™ Edit-R™ gene editing experiments
Safety data sheets
Selection guides
Technical manuals
-
CRISPR-Cas9 genome engineering with Cas9 nuclease expression plasmids and Edit-R synthetic RNA - Technical Manual
-
CRISPR-Cas9 genome engineering with Cas9 nuclease mRNA and Edit-R synthetic RNA - Technical Manual
-
CRISPR-Cas9 genome engineering with Cas9 nuclease protein and Edit-R synthetic guide RNA - Technical Manual
-
CRISPR-Cas9 genome engineering with lentiviral Cas9 particles and Edit-R synthetic guide RNA - Technical Manual
Related Products
A DNA-free option for Cas9 nuclease expression. Purified Cas9 nuclease protein with an enhanced hybrid NLS composition enabling more efficient nuclear delivery and gene editing. Suitable for a wide range of RNP workflows including nucleofection or co-transfection with Edit-R synthetic guide RNA for a completely DNA-free genome engineering system.
Purified Cas9 nuclease mRNA for co-transfection with synthetic guide RNA for a completely DNA-free genome engineering system
Choose from a range of popular cell line backgrounds, then simply deliver your CRISPR guide RNA for straightforward gene knockout
Positive control sgRNA for confirming gene editing with your experimental conditions, provided with or without primers for DNA mismatch detection assays
